Why it matters
- Replication and repair questions are high-yield because they let Step 1 test enzyme mechanism, disease association, pharmacology, and mutation biology in one vignette.
- The exam often asks you to choose between similar-sounding enzymes; one clue such as “needs an RNA primer” or “repairs thymine dimers” resolves the question fast.
- Many cancer predisposition syndromes make more sense once you know which repair pathway failed and what kind of DNA lesion accumulated.
- These concepts bridge directly into the next topics: replication sets up transcription template use and determines how genetic information reaches translation.
Key Terms & Must-Know Facts
1.1.1.1 Key terms
- Origin of replication: DNA sequence where replication begins
- Helicase: unwinds the double helix
- Single-strand binding proteins: stabilize separated DNA strands
- Primase: synthesizes short RNA primer
- DNA polymerase: extends DNA only in the \(5' \to 3'\) direction
- Leading strand: continuous synthesis toward fork movement
- Lagging strand: discontinuous synthesis away from fork movement
- Okazaki fragments: short DNA fragments on the lagging strand
- Ligase: seals phosphodiester backbone nicks
- Topoisomerase: relieves torsional strain ahead of the fork
- Proofreading: \(3' \to 5'\) exonuclease correction during synthesis
- Telomerase: reverse transcriptase that extends chromosome ends
- Nucleotide excision repair: removes bulky DNA lesions
- Base excision repair: removes small non-bulky damaged bases
- Mismatch repair: fixes replication misincorporation errors
1.1.1.2 Must-know facts
- DNA polymerases require a preexisting \(3'\)-OH; they cannot start de novo.
- Both leading and lagging strands are synthesized \(5' \to 3'\).
- Lagging-strand synthesis requires repeated priming.
- Proofreading is \(3' \to 5'\) exonuclease activity; primer removal in prokaryotes is \(5' \to 3'\) exonuclease by DNA polymerase I.
- Topoisomerase I creates single-strand cuts; topoisomerase II creates double-strand cuts.
- Fluoroquinolones inhibit bacterial topoisomerase II (DNA gyrase) and topoisomerase IV.
- Eukaryotic mRNA primers are removed differently from prokaryotic primer removal; Step 1 commonly tests the prokaryotic DNA polymerase I role.
- NER repairs thymine dimers caused by UV light.
- BER commonly repairs deamination damage.
- Mismatch repair defects are associated with hereditary nonpolyposis colorectal cancer.
- Double-strand break repair by homologous recombination is linked to BRCA tumor suppressor genes.
- Telomerase is active in germ cells, stem cells, and many cancers.
Exam takeaway: Use the order of enzymes to identify the single missing step in board-style replication questions.
flowchart LR A[Origin opens] --> B[Helicase unwinds DNA] B --> C[SSB proteins stabilize strands] C --> D[Primase lays RNA primer] D --> E[DNA polymerase extends 5 to 3] E --> F[Primer removal and replacement] F --> G[Ligase seals nick] E --> H[Proofreading reduces error rate] B --> I[Topoisomerase relieves supercoils]
Exam takeaway: Visualizing leading and lagging strands prevents confusion about strand direction and primer use.
Core content
1.1.1.3 Foundational replication mechanics: what enzyme acts next?
A Explanation
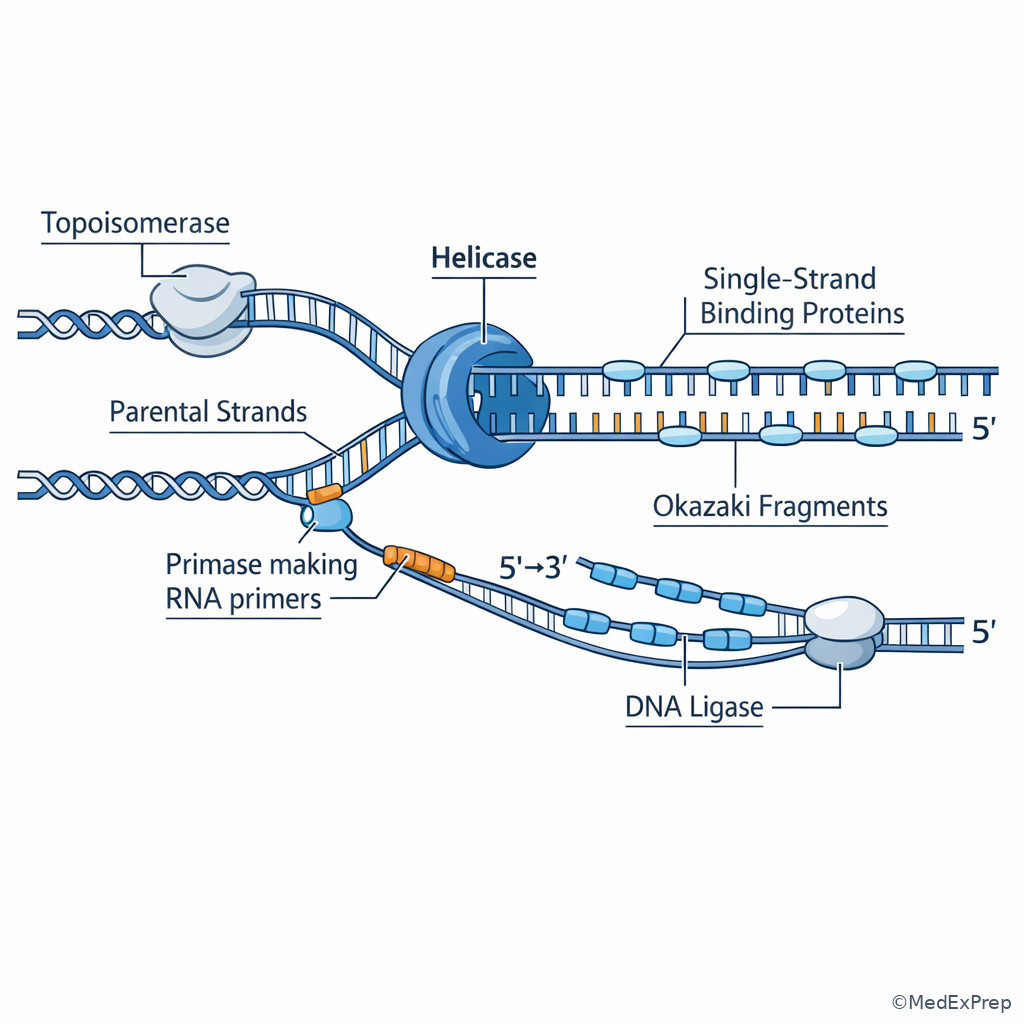
The main decision conflict in early replication questions is usually this: is the stem asking about unwinding, initiating, elongating, correcting, or sealing? The single rule that resolves most items is the sequence of events. At an origin of replication, helicase separates the strands, single-strand binding proteins prevent reannealing, primase creates a short RNA primer, and DNA polymerase extends from the primer using the template strand. If the stem says the enzyme starts synthesis de novo, choose primase; if it says the enzyme adds nucleotides to an existing 3' end, choose DNA polymerase; if it says the enzyme joins fragments, choose ligase. See Fig 1.
WHY THIS IS TESTED: The exam likes close-call wording such as “initiates nucleic acid synthesis,” “relieves supercoiling,” or “forms phosphodiester bonds,” because students often collapse these into one vague idea of “DNA replication.” Step 1 rewards knowing the unique job of each enzyme.
The highest-yield rule is directionality. DNA polymerases synthesize only \(5' \to 3'\), which means they read the template \(3' \to 5'\). Because the two parental strands are antiparallel, one daughter strand can be synthesized continuously toward fork progression (leading strand), while the other must be synthesized discontinuously as Okazaki fragments (lagging strand). If you see repeated RNA primers or fragment joining, the stem is signaling lagging-strand synthesis rather than leading-strand synthesis.
B Worked example
A 7-year-old boy is being evaluated for developmental delay and recurrent otitis media. During a research protocol, fibroblasts from his skin biopsy are exposed to a toxin that prevents formation of short RNA segments required before DNA elongation. His CBC and serum uric acid are normal, and his mother mentions he recently had a mild sunburn after summer camp. Which enzyme activity is most directly inhibited?
Reasoning chain: Identify task: this is asking which enzyme creates the short RNA segments required before DNA elongation. Extract key facts: “short RNA segments” and “required before DNA elongation” point to primers, not transcriptional mRNA production. Apply rule: DNA polymerase cannot start de novo and needs primase to create the primer. Eliminate distractors: helicase unwinds; ligase seals nicks; RNA polymerase II makes mRNA, not replication primers. Verify: the sunburn detail is noise and is not enough to make this a repair question.
C Exam trap
D Checkpoint
Question: A toxin selectively prevents formation of phosphodiester bonds between adjacent Okazaki fragments without affecting nucleotide addition to the growing chain. Which enzyme is most directly inhibited?
- DNA ligase — tempting because fragment joining is a late replication step; correct because sealing backbone nicks between Okazaki fragments is ligase’s defining job.
- DNA polymerase — tempting because polymerase forms phosphodiester bonds during chain elongation; wrong because the stem says nucleotide addition itself is intact and the defect is specifically between completed fragments.
- Primase — tempting because lagging-strand synthesis needs repeated priming; wrong because lack of primers would prevent fragment initiation, not leave completed fragments unsealed.
- Helicase — tempting because unwinding is required for all replication; wrong because helicase defects stop fork progression, not selective ligation of adjacent fragments.
1.1.1.4 Directionality, leading vs lagging strands, and proofreading
A Explanation
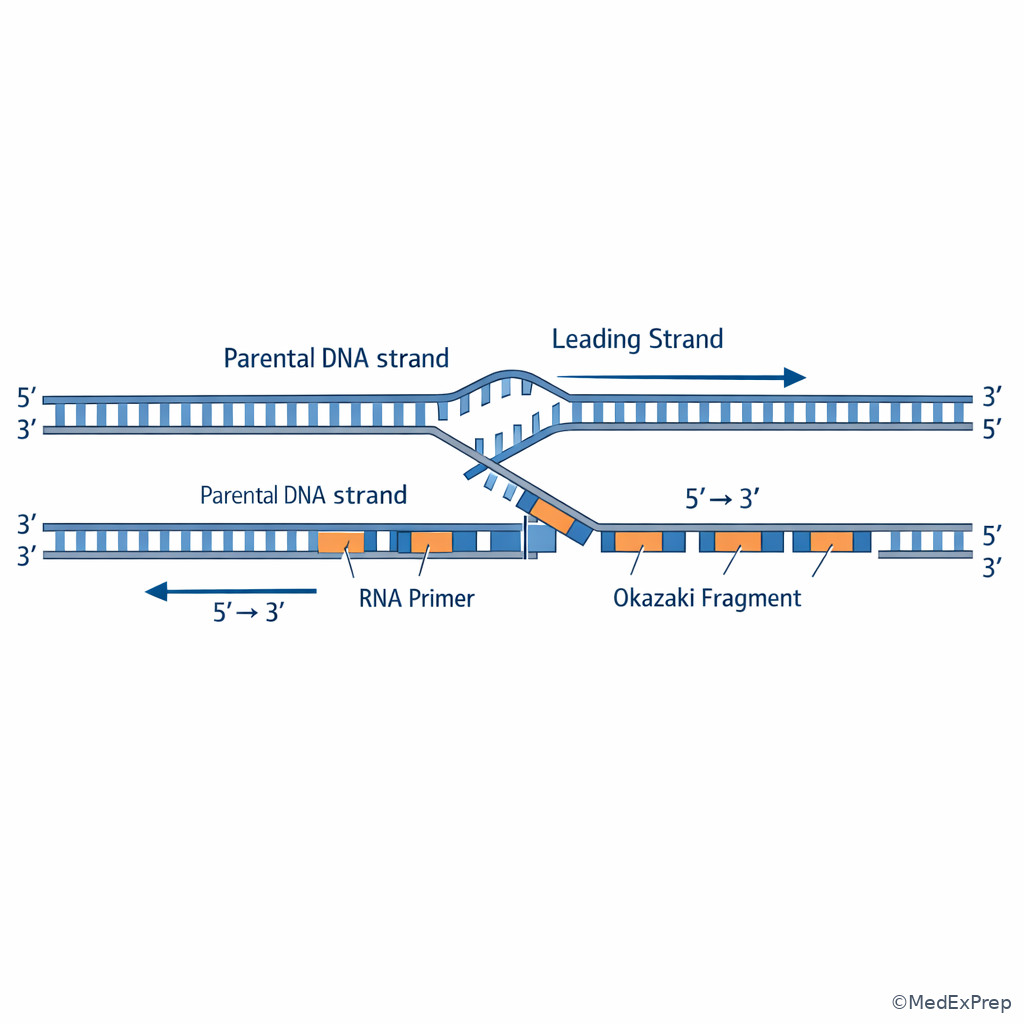
The decision conflict here is usually continuous versus discontinuous synthesis, or polymerization versus error correction. The single rule that resolves both is that all DNA polymerases synthesize DNA only \(5' \to 3'\). Because the template strands are antiparallel, the strand oriented to permit continuous synthesis toward fork movement becomes the leading strand. The opposite daughter strand is synthesized discontinuously as Okazaki fragments on the lagging strand. See Fig 2.
Proofreading is another favorite distinction. If the stem asks about correcting an incorrect base during active synthesis, choose a polymerase with \(3' \to 5'\) exonuclease activity. If the stem instead asks about removing an RNA primer in prokaryotes, the high-yield answer is DNA polymerase I with \(5' \to 3'\) exonuclease activity. Step 1 often pairs these two exonuclease directions in the same option set.
WHY THIS IS TESTED: Students remember “DNA goes \(5' \to 3'\)” but forget how that forces discontinuous lagging-strand synthesis and repeated primer placement. Once you know the geometry, many one-step mechanism questions become automatic.
Exam takeaway: Repeated primers and Okazaki fragments identify lagging-strand synthesis even when the stem hides the word “lagging.”
flowchart TD
A[DNA polymerase works only 5 to 3] --> B{Template orientation at fork}
B -->|Allows continuous extension| C[Leading strand]
B -->|Forces synthesis away from fork| D[Lagging strand]
D --> E[Repeated RNA primers]
E --> F[Okazaki fragments]
F --> G[Ligase seals fragments]
C --> H[Single primer usually sufficient] Exam takeaway: A labeled fork image helps separate continuous synthesis from proofreading and ligation tasks.
B Worked example
A 63-year-old man with hypertension and gout participates in a study of bacterial DNA replication. A mutant organism shows normal helicase activity and normal primer synthesis, but newly synthesized DNA contains many uncorrected base-pairing errors despite preserved RNA primer removal. Colony growth is slowed, and an intern incorrectly attributes this to a ligase defect. Which lost activity best explains the mutation burden?
Reasoning chain: Identify task: uncorrected base-pairing errors during synthesis. Extract key facts: helicase normal, primers normal, primer removal preserved. Apply rule: proofreading during synthesis is \(3' \to 5'\) exonuclease activity. Eliminate distractors: ligase seals nicks; \(5' \to 3'\) exonuclease removes primers; primase starts synthesis. Verify: the gout history is noise.
C Exam trap
D Checkpoint
Question: Which feature most strongly identifies a newly synthesized daughter strand as the lagging strand?
- Continuous extension toward the replication fork — tempting because all daughter strands are made during fork progression; wrong because this describes the leading strand.
- Requirement for repeated RNA primers — tempting because both strands need priming at some point; correct because only lagging-strand synthesis repeatedly re-initiates due to discontinuous fragment production.
- Use of DNA polymerase for nucleotide addition — tempting because polymerase is central to replication; wrong because both strands use DNA polymerase.
- Need for single-strand binding proteins — tempting because exposed DNA requires stabilization; wrong because both separated template strands need stabilization.
- The strand synthesized discontinuously is the lagging strand.
- True or False: DNA polymerase synthesizes DNA \(3' \to 5'\). False.
- Name the 2 high-yield exonuclease directions: \(3' \to 5'\) proofreading; \(5' \to 3'\) primer removal in prokaryotic DNA polymerase I.
1.1.1.5 Prokaryotic versus eukaryotic replication: which system is being tested?
A Explanation
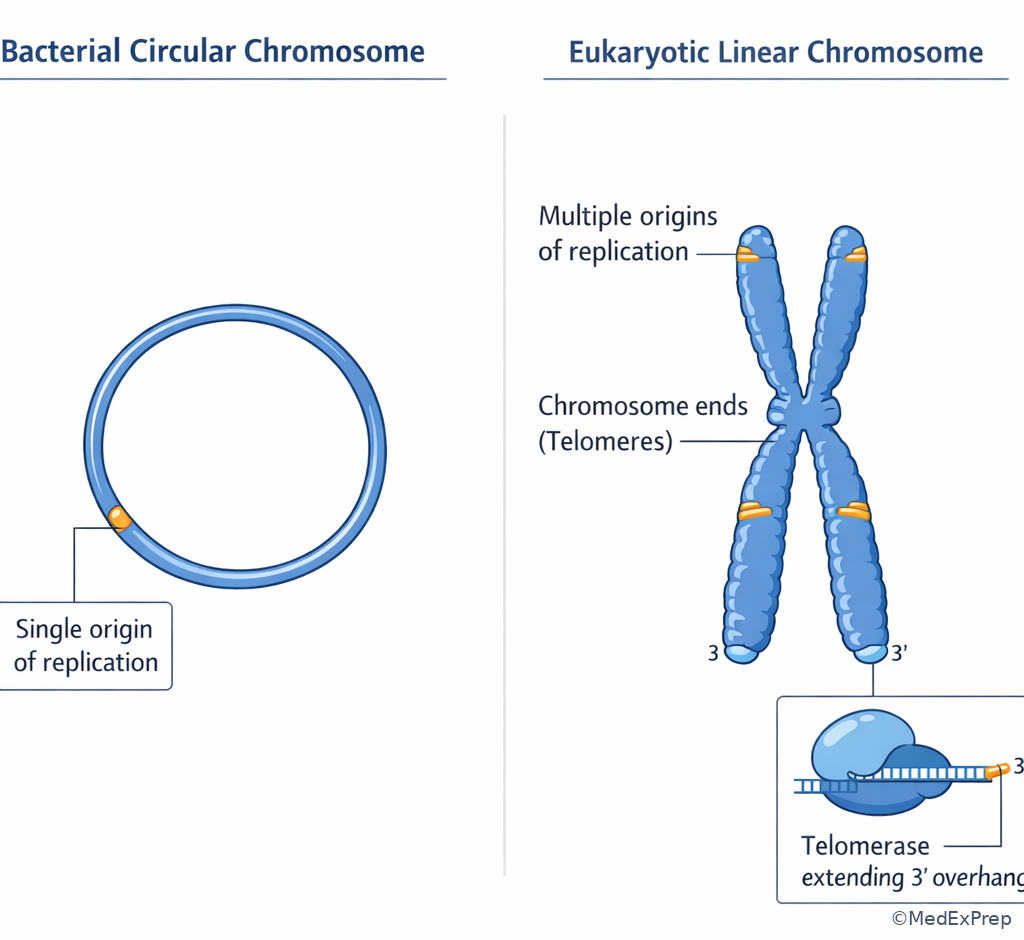
The decision conflict here is not “what happens in replication?” but which organism’s machinery does the question describe? The resolving clue is usually one of the following: multiple origins of replication, telomeres, histones, or named polymerase functions. Eukaryotes have multiple origins per chromosome and linear chromosomes that create an end-replication problem solved by telomerase. Prokaryotes typically have a single circular chromosome with one origin and no telomeres.
High-yield polymerase distinctions: in prokaryotes, DNA polymerase III is the main replicative enzyme and DNA polymerase I removes RNA primers and fills in DNA. In eukaryotes, Step 1 more often tests the broader idea that multiple nuclear polymerases exist and that primer removal is not simply “DNA polymerase I,” which is a prokaryotic detail. If a stem says a rapidly dividing malignant cell maintains chromosome ends and avoids senescence, think telomerase, a reverse transcriptase that carries its own RNA template. See Fig 3.
WHY THIS IS TESTED: Board questions often bait students with a true fact from one domain placed into the wrong biological setting, such as using DNA polymerase I in a eukaryotic stem or asking about telomerase in bacteria.
Exam takeaway: Distinguish organism type first when options mix bacterial polymerases with telomerase or multiple origins.
flowchart LR
A[Replication stem] --> B{Key clue}
B -->|Single circular chromosome| C[Think prokaryote]
B -->|Multiple origins on linear chromosomes| D[Think eukaryote]
C --> E[DNA pol III extends DNA]
C --> F[DNA pol I removes RNA primer]
D --> G[Telomeres create end-replication problem]
D --> H[Telomerase extends chromosome ends] Exam takeaway: A telomere image makes the end-replication problem easier to recall than text alone.
B Worked example
A 52-year-old woman with myelodysplastic syndrome and type 2 diabetes enrolls in a cancer genomics study. Tumor cells from her bone marrow demonstrate maintenance of chromosome ends despite repeated divisions. A classmate suggests DNA polymerase I is responsible because the cells divide rapidly. Which enzyme best explains the observed finding?
Reasoning chain: Identify task: maintenance of chromosome ends in repeatedly dividing eukaryotic cells. Extract key facts: tumor cells, chromosome ends, repeated divisions. Apply rule: telomerase is a reverse transcriptase that extends telomeres. Eliminate distractors: DNA polymerase I is a prokaryotic primer-removal enzyme; ligase seals nicks; helicase unwinds but does not solve end shortening. Verify: the marrow malignancy detail supports high cellular turnover but the specific clue is chromosome-end maintenance.
C Exam trap
D Checkpoint
Question: Which finding most strongly supports that a replication vignette is describing a eukaryotic cell rather than a prokaryote?
- Need for RNA primer formation — tempting because primers are essential in replication; wrong because both systems require primers.
- DNA synthesis in the \(5' \to 3'\) direction — tempting because directionality is central; wrong because both prokaryotes and eukaryotes synthesize DNA this way.
- Presence of multiple origins on a linear chromosome — tempting because it sounds detailed; correct because this is a classic eukaryotic replication feature.
- Use of helicase to separate strands — tempting because unwinding is fundamental; wrong because both systems use helicase function.
1.1.1.6 DNA damage and repair pathways: which lesion goes with which mechanism?
A Explanation
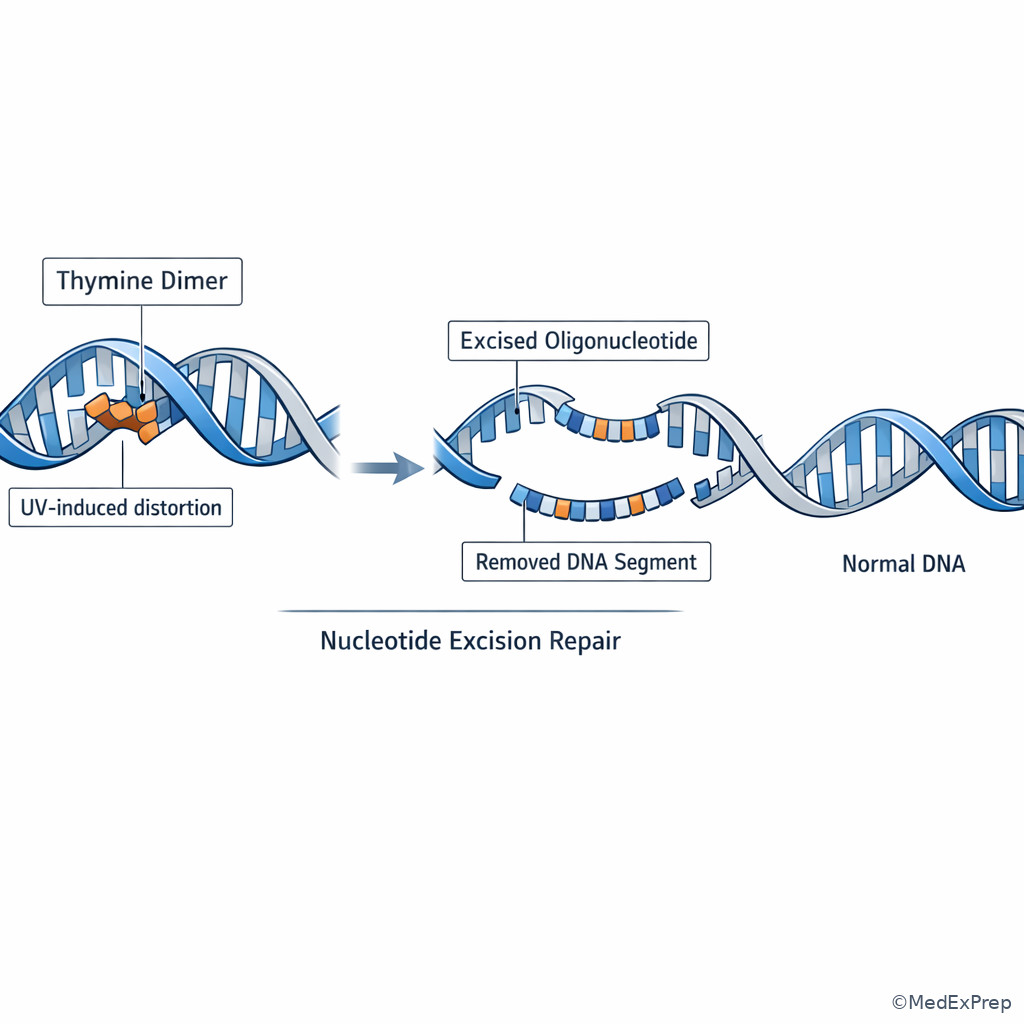
The decision conflict here is small-base damage, bulky helix distortion, replication mismatch, or double-strand break? The single resolving clue is the type of DNA lesion. If the stem says thymine dimers or another bulky helix-distorting lesion, choose nucleotide excision repair (NER). If it says deaminated, oxidized, or alkylated base with no major helix distortion, choose base excision repair (BER). If it says replication error that escaped proofreading, choose mismatch repair (MMR). If it says double-strand breaks, think homologous recombination or nonhomologous end joining.
NER is the high-yield answer to UV-light questions because UV commonly creates pyrimidine dimers that distort the helix. BER removes the abnormal base with a DNA glycosylase, then repairs the site. MMR functions after replication and is especially tied to microsatellite instability when defective. See Fig 4.
WHY THIS IS TESTED: Repair pathways are one of the cleanest ways for Step 1 to ask mechanism-based questions. The stem often gives one lesion type and then asks for the disease, enzyme class, or cancer syndrome associated with failure of that pathway.
Exam takeaway: Match the lesion first, then choose the repair pathway; most repair questions become one-step decisions.
flowchart TD
A[DNA damage identified] --> B{What kind of lesion?}
B -->|Bulky helix distortion or thymine dimer| C[Nucleotide excision repair]
B -->|Single damaged base such as deamination| D[Base excision repair]
B -->|Replication mismatch| E[Mismatch repair]
B -->|Double-strand break| F[Homologous recombination or NHEJ]
C --> G[Xeroderma pigmentosum if defective]
E --> H[Lynch syndrome if defective]
F --> I[BRCA association with homologous recombination defects] Exam takeaway: Seeing the helix distortion helps distinguish nucleotide excision repair from repair of single damaged bases.
B Worked example
An 11-year-old boy with fair skin, numerous freckles, and a history of recurrent blistering sunburns develops a scaly lesion on the nose. He also has mild asthma and seasonal allergies; CBC and renal function are normal. Skin cells show impaired removal of ultraviolet-induced DNA damage that distorts the helix. Which repair pathway is most directly defective?
Reasoning chain: Identify task: repair pathway for UV-induced helix-distorting damage. Extract key facts: severe sun sensitivity, skin lesion, UV-induced distortion. Apply rule: thymine dimers are bulky lesions repaired by nucleotide excision repair. Eliminate distractors: BER handles small base modifications; MMR handles replication mismatches; homologous recombination fixes double-strand breaks. Verify: fair skin and freckles support xeroderma pigmentosum rather than being mere cosmetic noise.
C Exam trap
D Checkpoint
Question: A defect in which repair pathway most directly leads to persistence of replication mismatches and microsatellite instability?
- Nucleotide excision repair — tempting because it removes damaged DNA segments; wrong because it targets bulky lesions such as thymine dimers rather than post-replication base mismatches.
- Base excision repair — tempting because it handles abnormal bases; wrong because BER addresses chemically altered individual bases, not insertion-deletion mismatches from replication.
- Mismatch repair — tempting because the name mirrors the lesion; correct because it corrects mispaired nucleotides that escaped polymerase proofreading.
- Nonhomologous end joining — tempting because DNA breaks can destabilize the genome; wrong because this pathway repairs double-strand breaks, not replication mismatches.
- The repair pathway for thymine dimers is nucleotide excision repair.
- True or False: mismatch repair fixes UV-induced pyrimidine dimers. False.
- Name the 3 major lesion-pathway pairs: bulky lesion–NER, single-base damage–BER, replication mismatch–MMR.
1.1.1.7 Repair defects, cancer syndromes, and pharmacologic/testable links
A Explanation
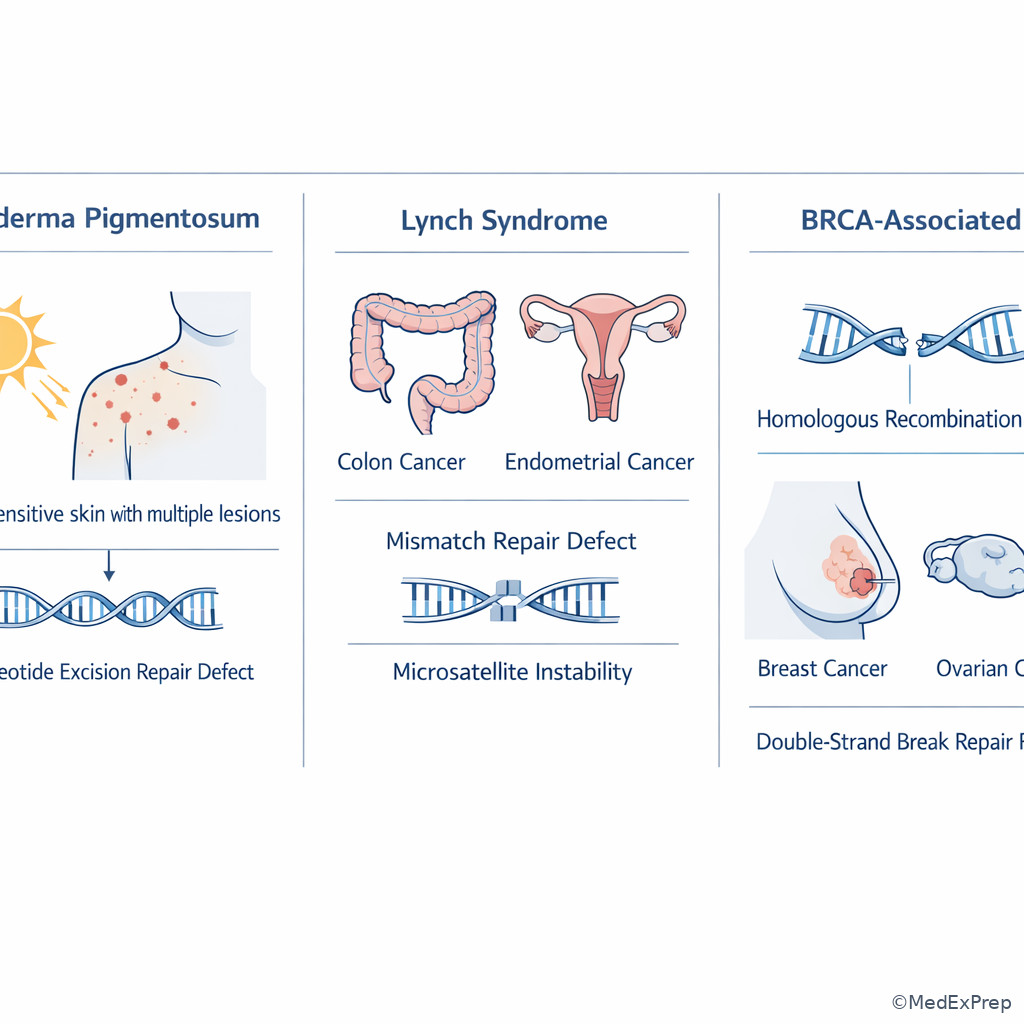
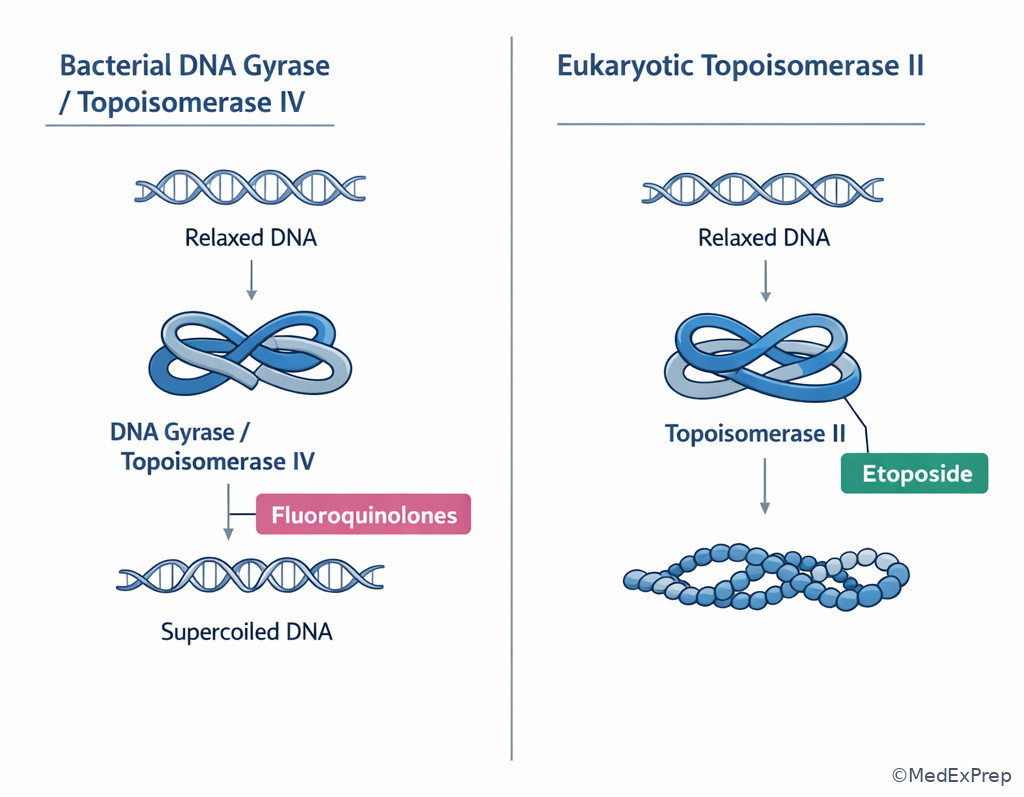
The integration-level decision conflict is usually which syndrome fits the repair defect or which drug targets a related replication enzyme. The resolving clue is often a single associated phenotype. Xeroderma pigmentosum points to defective NER and inability to repair UV-induced thymine dimers. Lynch syndrome points to defective mismatch repair with microsatellite instability. BRCA1/BRCA2 defects impair homologous recombination for double-strand break repair, predisposing to breast and ovarian cancers. At the pharmacology interface, fluoroquinolones inhibit bacterial topoisomerase II (DNA gyrase) and topoisomerase IV; etoposide inhibits eukaryotic topoisomerase II.
Another classic board angle is telomerase and malignant immortality. Many tumor cells reactivate telomerase, allowing repeated division without critical telomere shortening. This is not the same as a repair pathway defect, but it belongs in the same “genome stability” family of ideas. See Fig 5.
WHY THIS IS TESTED: These are integrated, high-yield pattern-recognition questions. The exam can give you a disease phenotype, a lesion type, a tumor predisposition, or a drug mechanism and ask for the missing molecular step.
Exam takeaway: Match a classic phenotype to its failed pathway before considering broader cancer associations.
Exam takeaway: Distinguishing bacterial from eukaryotic topoisomerase targets helps separate antibiotic from chemotherapy mechanisms.
B Worked example
A 34-year-old woman with a strong family history of breast and ovarian cancer is found to have a germline mutation in a tumor suppressor gene involved in high-fidelity repair of DNA double-strand breaks. She also has hypothyroidism and iron-deficiency anemia after a recent pregnancy. Which DNA repair process is most directly impaired?
Reasoning chain: Identify task: repair process impaired by inherited breast/ovarian cancer predisposition. Extract key facts: germline tumor suppressor mutation, double-strand break repair, family history of breast and ovarian cancer. Apply rule: BRCA genes mediate homologous recombination repair. Eliminate distractors: NER is for thymine dimers; BER for damaged single bases; MMR for replication mismatches and Lynch syndrome. Verify: postpartum anemia and hypothyroidism are distractors with no bearing on repair pathway selection.
C Exam trap
D Checkpoint
Question: A drug inhibits bacterial DNA gyrase, leading to impaired supercoil relaxation during replication. Which medication class works by this mechanism?
- Fluoroquinolones — tempting because they are antibacterial agents with DNA synthesis effects; correct because they inhibit bacterial topoisomerase II (DNA gyrase) and topoisomerase IV.
- Rifamycins — tempting because they target nucleic acid processes in bacteria; wrong because they inhibit DNA-dependent RNA polymerase, affecting transcription rather than DNA gyrase.
- Tetracyclines — tempting because they inhibit bacterial growth; wrong because they target the 30S ribosomal subunit and therefore translation, not DNA supercoiling.
- Sulfonamides — tempting because they impair bacterial replication indirectly; wrong because they inhibit folate synthesis rather than topoisomerase function.
Exam Traps & Differentiators
- Primase vs RNA polymerase II — tempting because both synthesize RNA; eliminate with the clue “short primer for DNA replication” versus “mRNA transcription.”
- Proofreading vs mismatch repair — tempting because both fix wrong bases; eliminate with timing: proofreading happens during synthesis, mismatch repair acts after replication errors escape polymerase correction.
- BER vs NER — tempting because both include “excision”; eliminate with lesion type: BER removes a single altered base, NER removes bulky helix-distorting lesions.
- DNA polymerase I vs telomerase — tempting because both fill DNA-related gaps; eliminate with chromosome-end language, which points to telomerase in eukaryotes.
- Fluoroquinolone vs rifampin — tempting because both target nucleic-acid-associated enzymes in bacteria; eliminate with “supercoiling/topoisomerase” versus “RNA polymerase/transcription.”
Most common wrong answer and why: Nucleotide excision repair questions are frequently missed because learners choose the more familiar phrase “base excision repair” whenever they see DNA damage. That error happens when they classify by the word “damage” instead of by the lesion shape. Bulky, helix-distorting UV damage belongs to NER.
1.1.1.8 Looks similar but isn't
| Confusable pair | Why both seem plausible | Single discriminating clue | Choose |
|---|---|---|---|
| Primase vs RNA polymerase II | Both synthesize RNA | Replication primer at fork | Primase |
| Proofreading vs mismatch repair | Both correct mispaired bases | Error corrected during synthesis vs after replication | Proofreading for during synthesis; mismatch repair for post-replication |
| NER vs BER | Both are excision pathways | Bulky helix distortion or thymine dimers | NER |
| DNA pol I vs ligase | Both function late on lagging strand | Primer removal/fill-in versus nick sealing | DNA pol I for primer replacement; ligase for final sealing |
| Fluoroquinolone vs rifampin | Both impair bacterial nucleic acid handling | Supercoil relief versus RNA synthesis | Fluoroquinolone if topoisomerase is named |
If the stem says...
- “Requires a free 3'-OH” → think DNA polymerase cannot initiate de novo.
- “Repeated short RNA segments” → think lagging strand and primase.
- “Ultraviolet-induced thymine dimers” → think nucleotide excision repair.
- “Microsatellite instability” → think mismatch repair defect.
- “Maintenance of chromosome ends in cancer cells” → think telomerase.
- “Supercoiling ahead of the replication fork” → think topoisomerase.
Tables
| Replication component | Main function | Board clue | Classic distractor to reject |
|---|---|---|---|
| Helicase | Separates DNA strands | Unwinds double helix at fork | Topoisomerase if the stem says upstream torsional strain |
| Topoisomerase | Relieves supercoiling | Cuts and reseals DNA ahead of fork | Helicase if the stem says strand separation |
| SSB proteins | Stabilize single strands | Prevent reannealing | Helicase if the stem asks who actually opens the helix |
| Primase | Makes RNA primer | Starts synthesis de novo | RNA polymerase II in eukaryotic transcription questions |
| DNA polymerase | Adds nucleotides to 3'-OH | Extends DNA 5' to 3' | Ligase if the defect is between fragments rather than along chain growth |
| DNA polymerase I (prokaryotes) | Removes RNA primer and fills gap | \(5' \to 3'\) exonuclease | Proofreading if the stem is about mispaired bases during synthesis |
| Ligase | Seals nicks | Joins Okazaki fragments | Polymerase if nucleotide addition is the actual defect |
| Telomerase | Extends chromosome ends | Reverse transcriptase in germ cells and cancers | DNA pol I or ligase in eukaryotic chromosome-end questions |
| Comparison feature | Leading strand | Lagging strand |
|---|---|---|
| Direction of synthesis | \(5' \to 3'\) | \(5' \to 3'\) |
| Relation to fork movement | Toward fork progression | Away from fork progression |
| Continuity | Continuous | Discontinuous |
| Primer requirement | Initial primer | Repeated primers |
| Fragment formation | No Okazaki fragments | Okazaki fragments present |
| Need for ligase joining | Minimal relative role | Essential to seal fragment junctions |
| Lesion / clinical clue | Repair pathway | Associated disorder | High-yield discriminator |
|---|---|---|---|
| UV-induced thymine dimers | Nucleotide excision repair | Xeroderma pigmentosum | Bulky helix distortion |
| Deaminated or oxidized base | Base excision repair | No single classic syndrome tested as often as XP/Lynch | Single abnormal base, not bulky lesion |
| Replication mismatch, microsatellite instability | Mismatch repair | Lynch syndrome | Post-replication error correction |
| Double-strand breaks | Homologous recombination / NHEJ | BRCA-associated cancers for homologous recombination defects | High-fidelity repair using homologous template |
| Shortened chromosome ends in replicating eukaryotic cells | Telomerase-related maintenance | Cellular senescence if absent; many cancers if activated | Chromosome-end problem, not classic lesion excision |
| Drug / target category | Target | System | Why it is confusable |
|---|---|---|---|
| Fluoroquinolones | Bacterial topoisomerase II and IV | Replication/supercoiling | May be confused with transcription inhibitors because both affect nucleic acids |
| Rifamycins | DNA-dependent RNA polymerase | Transcription | Also a nucleic acid drug but does not target replication fork mechanics |
| Etoposide | Eukaryotic topoisomerase II | Antineoplastic DNA handling | Shares topoisomerase target family with fluoroquinolones but in a different system |
| Tetracyclines | 30S ribosomal subunit | Translation | Can be chosen incorrectly if the student notices “bacterial growth inhibition” but not the actual mechanism |
Algorithm / Approach
Exam takeaway: Start by classifying the task as normal replication, lesion repair, or syndrome association before choosing an enzyme.
Use this flow on exam day. The highest-value move is to avoid premature answer selection. First decide whether the question is about normal synthesis, repair after damage, or syndrome/pharmacology integration. Then use one specific clue to separate close distractors. See Fig 6.
Rapid Review
- Primase → initiates replication with RNA primer; RNA polymerase II does not run the fork
- DNA polymerase → needs preexisting \(3'\)-OH; cannot start de novo
- Lagging strand → repeated primers and Okazaki fragments
- Ligase → seals nicks after fragment synthesis, not during primer creation
- Proofreading → \(3' \to 5'\) exonuclease during synthesis
- DNA pol I → \(5' \to 3'\) exonuclease removes RNA primers in prokaryotes
- Topoisomerase → relieves supercoils ahead of fork; helicase separates strands at fork
- Telomerase → reverse transcriptase that extends chromosome ends in eukaryotes
- NER → bulky lesions, especially UV thymine dimers
- BER → single damaged base such as deamination
- MMR → post-replication mismatch correction; Lynch association
- Homologous recombination → high-fidelity double-strand break repair; BRCA link
- Xeroderma pigmentosum → severe photosensitivity from defective NER
- Fluoroquinolones → bacterial topoisomerase inhibition, not ribosomal inhibition
Self-check quiz
1.1.1.9 Question 1
A 19-year-old man in a cell biology lab is exposed to a compound that prevents the enzyme responsible for initiating DNA synthesis by making a short RNA segment. He has mild allergic rhinitis and a normal CBC. Which enzyme is most directly inhibited?
- DNA ligase
- Primase
- Topoisomerase II
- DNA polymerase
1.1.1.10 Question 2
A bacterial mutant has impaired removal of RNA primers from newly synthesized DNA but retains normal elongation by the main replicative polymerase. Which enzyme is most directly defective?
- DNA polymerase I
- DNA polymerase III
- Helicase
- Ligase
1.1.1.11 Question 3
A 9-year-old boy with fair skin, multiple freckles, and recurrent blistering after minimal sun exposure is found to have an early basal cell carcinoma. He also has asthma and a recent normal urinalysis. The most likely underlying defect impairs repair of which type of DNA lesion?
- Replication mismatches after polymerase slippage
- Bulky ultraviolet-induced pyrimidine dimers
- Single cytosine deamination events without helix distortion
- Double-strand breaks repaired using a sister chromatid
1.1.1.12 Question 4
A 46-year-old woman with obesity and iron-deficiency anemia has colon cancer; her mother and maternal aunt both had endometrial cancer. Tumor testing shows microsatellite instability. Which DNA repair pathway is most likely defective?
- Base excision repair
- Nucleotide excision repair
- Mismatch repair
- Homologous recombination
1.1.1.13 Question 5
A 67-year-old man with COPD and type 2 diabetes develops pneumonia due to a gram-negative organism. He is treated with an antibiotic that inhibits bacterial DNA gyrase, impairing relaxation of supercoiled DNA during replication. Which drug class was most likely used?
- Fluoroquinolones
- Macrolides
- Rifamycins
- Sulfonamides
1.1.1.14 Question 6
A 32-year-old woman with a strong family history of breast and ovarian cancer is found to carry a germline mutation in a tumor suppressor gene. Her cells show reduced ability to repair DNA double-strand breaks accurately after ionizing radiation exposure. She also has migraine headaches and a mildly elevated LDL level. Which process is most directly impaired?
- Mismatch repair
- Homologous recombination
- Nucleotide excision repair
- Base excision repair
1.1.1.15 Question 7
A 58-year-old man with myelofibrosis undergoes tumor sequencing. The malignant clone demonstrates persistent maintenance of chromosome ends, allowing many additional rounds of cell division. He also has hyperuricemia and splenomegaly. Which enzyme activity best explains this finding?
- DNA-dependent RNA polymerase
- DNA polymerase I \(5' \to 3'\) exonuclease
- Reverse transcriptase that extends telomeric repeats
- DNA ligase sealing phosphodiester nicks
1.1.1.16 Question 8
A researcher tracks DNA synthesis at a replication fork in human cells. One daughter strand is synthesized in short fragments requiring repeated initiation events, while the other is synthesized continuously. Which single feature best explains why discontinuous synthesis occurs on only one strand?
- DNA ligase acts only on one of the two daughter strands
- Helicase unwinds only one parental strand at a time
- DNA polymerase can extend DNA only in the \(5' \to 3'\) direction
- Topoisomerase creates transient breaks in only one parental strand
Answer key
1.1.1.17 Question 1 — Correct answer: B. Primase
Why B is correct: Primase synthesizes the short RNA primer required to initiate DNA replication. DNA polymerase needs a preexisting \(3'\)-OH and cannot begin de novo. This is the classic replication-initiation clue. Sources: First Aid for the USMLE Step 1; Alberts et al., Molecular Biology of the Cell.
- A. DNA ligase — tempting because it is essential in replication; incorrect because ligase joins nicks after DNA fragments are already formed. Discriminating clue: “initiating DNA synthesis by making a short RNA segment.”
- B. Primase — tempting and correct because it is the enzyme that creates the RNA primer needed before elongation can start. Discriminating clue: “short RNA segment.”
- C. Topoisomerase II — tempting because it also acts early during replication; incorrect because it relieves supercoiling rather than synthesizing a primer. Discriminating clue: the stem focuses on RNA initiation, not torsional strain.
- D. DNA polymerase — tempting because it is the main DNA-synthesizing enzyme; incorrect because it extends from an existing primer rather than creating one. Discriminating clue: “initiating” rather than “elongating.”
1.1.1.18 Question 2 — Correct answer: A. DNA polymerase I
Why A is correct: In prokaryotes, DNA polymerase I removes RNA primers using \(5' \to 3'\) exonuclease activity and fills the resulting gap with DNA. DNA polymerase III is the main replicative polymerase but does not perform the classic high-yield primer removal role. Sources: First Aid for the USMLE Step 1; Murray et al., Harper’s Illustrated Biochemistry.
- A. DNA polymerase I — tempting and correct because primer removal in bacteria is its signature tested function. Discriminating clue: “bacterial mutant” with impaired RNA primer removal.
- B. DNA polymerase III — tempting because it is the major bacterial DNA polymerase; incorrect because elongation by the main replicative polymerase is said to remain normal. Discriminating clue: “retains normal elongation by the main replicative polymerase.”
- C. Helicase — tempting because unwinding is required for replication; incorrect because helicase does not remove RNA primers. Discriminating clue: the defect occurs after primers are already present.
- D. Ligase — tempting because it acts after primer replacement on the lagging strand; incorrect because ligase seals nicks and does not excise RNA primers. Discriminating clue: “removal of RNA primers.”
1.1.1.19 Question 3 — Correct answer: B. Bulky ultraviolet-induced pyrimidine dimers
Why B is correct: Severe photosensitivity with early skin cancer suggests xeroderma pigmentosum, caused by defective nucleotide excision repair. NER removes bulky helix-distorting lesions, especially UV-induced pyrimidine dimers. Sources: Robbins & Cotran, Pathologic Basis of Disease; First Aid for the USMLE Step 1.
- A. Replication mismatches after polymerase slippage — tempting because DNA repair defects can predispose to cancer; incorrect because this points to mismatch repair and Lynch syndrome, not extreme photosensitivity. Discriminating clue: “recurrent blistering after minimal sun exposure.”
- B. Bulky ultraviolet-induced pyrimidine dimers — tempting and correct because UV sensitivity with skin malignancy is classic for NER failure. Discriminating clue: sun sensitivity plus early skin cancer.
- C. Single cytosine deamination events without helix distortion — tempting because altered bases also require repair; incorrect because these small lesions are handled by BER and do not produce the classic XP phenotype. Discriminating clue: ultraviolet exposure rather than spontaneous base damage.
- D. Double-strand breaks repaired using a sister chromatid — tempting because defective repair can cause cancer predisposition; incorrect because this describes homologous recombination defects such as BRCA-associated disease, not marked childhood photosensitivity. Discriminating clue: cutaneous UV phenotype rather than breast/ovarian cancer clustering.
1.1.1.20 Question 4 — Correct answer: C. Mismatch repair
Why C is correct: Colon cancer plus a family history of endometrial cancer and microsatellite instability strongly indicates Lynch syndrome, which results from mismatch repair defects. MMR corrects replication errors that escaped polymerase proofreading. Sources: Robbins & Cotran; Pathoma; First Aid for the USMLE Step 1.
- A. Base excision repair — tempting because it also corrects DNA abnormalities; incorrect because BER repairs small non-bulky base lesions, not microsatellite instability. Discriminating clue: “microsatellite instability.”
- B. Nucleotide excision repair — tempting because “excision repair” sounds broadly relevant to cancer; incorrect because NER is tied to bulky lesions and xeroderma pigmentosum, not Lynch syndrome. Discriminating clue: colon/endometrial cancer family pattern rather than UV sensitivity.
- C. Mismatch repair — tempting and correct because post-replication mismatch defects cause microsatellite instability and Lynch syndrome. Discriminating clue: microsatellite instability with colon/endometrial cancer clustering.
- D. Homologous recombination — tempting because inherited DNA repair defects often lead to cancer; incorrect because homologous recombination defects classically suggest BRCA-related breast/ovarian cancer risk. Discriminating clue: the cancer pattern is Lynch, not BRCA.
1.1.1.21 Question 5 — Correct answer: A. Fluoroquinolones
Why A is correct: Fluoroquinolones inhibit bacterial DNA gyrase (topoisomerase II) and topoisomerase IV, impairing DNA supercoil relaxation and replication. This is a classic pharmacology crossover with replication enzymes. Sources: First Aid for the USMLE Step 1; Katzung, Basic & Clinical Pharmacology.
- A. Fluoroquinolones — tempting and correct because the stem names bacterial DNA gyrase directly. Discriminating clue: “inhibits bacterial DNA gyrase.”
- B. Macrolides — tempting because they are used for respiratory infections; incorrect because they inhibit the 50S ribosomal subunit and therefore translation. Discriminating clue: supercoiling, not ribosomal protein synthesis.
- C. Rifamycins — tempting because they target nucleic acid production; incorrect because they inhibit bacterial RNA polymerase, affecting transcription rather than gyrase. Discriminating clue: DNA gyrase rather than RNA polymerase.
- D. Sulfonamides — tempting because they impair bacterial growth through nucleotide depletion; incorrect because they inhibit folate synthesis upstream, not topoisomerase activity. Discriminating clue: direct enzyme target is DNA gyrase.
1.1.1.22 Question 6 — Correct answer: B. Homologous recombination
Why B is correct: BRCA tumor suppressor genes are required for high-fidelity homologous recombination repair of DNA double-strand breaks. A family history of breast and ovarian cancer plus impaired accurate repair after ionizing radiation strongly points to this pathway. Sources: Robbins & Cotran; Harrison’s Principles of Internal Medicine; First Aid for the USMLE Step 1.
- A. Mismatch repair — tempting because it is another inherited cancer-related repair pathway; incorrect because MMR defects are linked to Lynch syndrome and microsatellite instability rather than BRCA-pattern cancers. Discriminating clue: breast/ovarian family history plus double-strand break language.
- B. Homologous recombination — tempting and correct because BRCA proteins mediate accurate double-strand break repair using a homologous template. Discriminating clue: “repair DNA double-strand breaks accurately.”
- C. Nucleotide excision repair — tempting because it also prevents mutation accumulation; incorrect because it targets bulky lesions like UV-induced dimers, not ionizing-radiation-induced double-strand breaks. Discriminating clue: double-strand breaks rather than UV damage.
- D. Base excision repair — tempting because it repairs certain DNA injuries; incorrect because BER addresses isolated damaged bases rather than major double-strand breaks. Discriminating clue: need for accurate repair after ionizing radiation.
1.1.1.23 Question 7 — Correct answer: C. Reverse transcriptase that extends telomeric repeats
Why C is correct: Telomerase is a reverse transcriptase that extends telomeric DNA repeats using an internal RNA template, allowing continued cell division and replicative immortality in many cancers. Sources: First Aid for the USMLE Step 1; Alberts et al., Molecular Biology of the Cell.
- A. DNA-dependent RNA polymerase — tempting because tumors have active transcription; incorrect because transcription does not solve chromosome-end shortening. Discriminating clue: “maintenance of chromosome ends.”
- B. DNA polymerase I \(5' \to 3'\) exonuclease — tempting because it sounds DNA-replication related; incorrect because it is a prokaryotic primer-removal function and not relevant to telomeres in human tumor cells. Discriminating clue: human malignant clone with chromosome ends.
- C. Reverse transcriptase that extends telomeric repeats — tempting and correct because telomerase prevents replicative senescence by maintaining telomeres. Discriminating clue: persistent chromosome-end maintenance.
- D. DNA ligase sealing phosphodiester nicks — tempting because ligase supports DNA replication completion; incorrect because nick sealing does not address terminal DNA shortening. Discriminating clue: the problem is end maintenance, not fragment joining.
1.1.1.24 Question 8 — Correct answer: C. DNA polymerase can extend DNA only in the \(5' \to 3'\) direction
Why C is correct: Because DNA polymerase synthesizes only \(5' \to 3'\), one antiparallel daughter strand can be made continuously toward the fork, whereas the other must be synthesized discontinuously in fragments. This directionality is the fundamental reason for lagging-strand synthesis. Sources: Costanzo, Physiology; First Aid for the USMLE Step 1; Harper’s Illustrated Biochemistry.
- A. DNA ligase acts only on one of the two daughter strands — tempting because ligase is particularly important for Okazaki fragments; incorrect because lagging synthesis arises before ligase acts and is caused by polymerase directionality. Discriminating clue: asks why discontinuous synthesis occurs at all.
- B. Helicase unwinds only one parental strand at a time — tempting because helicase is central at the fork; incorrect because helicase separates the duplex but does not determine continuous versus discontinuous synthesis. Discriminating clue: strand asymmetry comes from synthesis direction, not unwinding order.
- C. DNA polymerase can extend DNA only in the \(5' \to 3'\) direction — tempting and correct because antiparallel templates plus one-way polymerase chemistry force lagging-strand fragmentation. Discriminating clue: one strand is continuous and the other fragmented.
- D. Topoisomerase creates transient breaks in only one parental strand — tempting because topoisomerases alter DNA structure during replication; incorrect because supercoil relief does not explain why only one daughter strand is discontinuous. Discriminating clue: repeated initiation events and short fragments indicate a directionality problem.
