Why it matters
- Replication and repair questions are classic “mechanism from phenotype” items: the stem gives photosensitivity, colon cancer risk, or chemotherapy toxicity, and you must identify the defective pathway.
- These pathways connect biochemistry, genetics, pathology, pharmacology, and oncology, making them highly reusable across organ systems.
- Many board distractors differ by one clue: UV damage versus deamination, mismatch after replication versus bulky adducts before transcription.
- Understanding repair logic also helps with upcoming transcription and translation topics, because unrepaired DNA lesions alter RNA synthesis and downstream protein production.
Key Terms & Must-Know Facts
1.1.1.1 Key terms
- Origin of replication: DNA sequence where replication begins
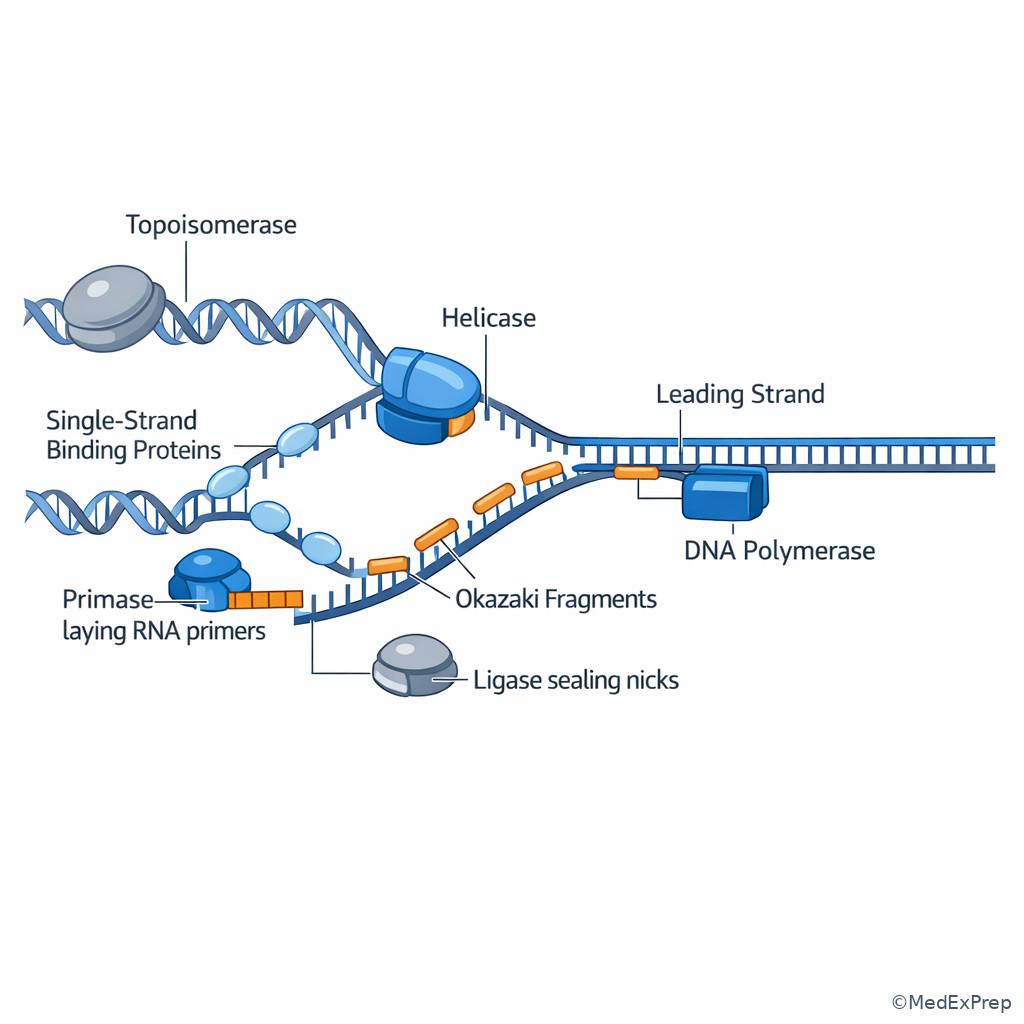
- Replication fork: Y-shaped region where parental strands separate
- Helicase: Enzyme that unwinds double-stranded DNA
- Single-strand binding proteins: Stabilize separated DNA strands
- Topoisomerase: Relieves torsional strain ahead of the fork
- Primase: Synthesizes short RNA primer
- DNA polymerase: Adds nucleotides to the 3' end of a growing strand
- Leading strand: Continuously synthesized strand
- Lagging strand: Discontinuously synthesized strand
- Okazaki fragments: Short DNA segments on the lagging strand
- Ligase: Seals phosphodiester backbone breaks
- Proofreading: \(3' \to 5'\) exonuclease correction during synthesis
- Mismatch repair: Post-replication correction of base-pairing errors
- Base excision repair: Removal of small non-bulky lesions
- Nucleotide excision repair: Removal of bulky helix-distorting lesions
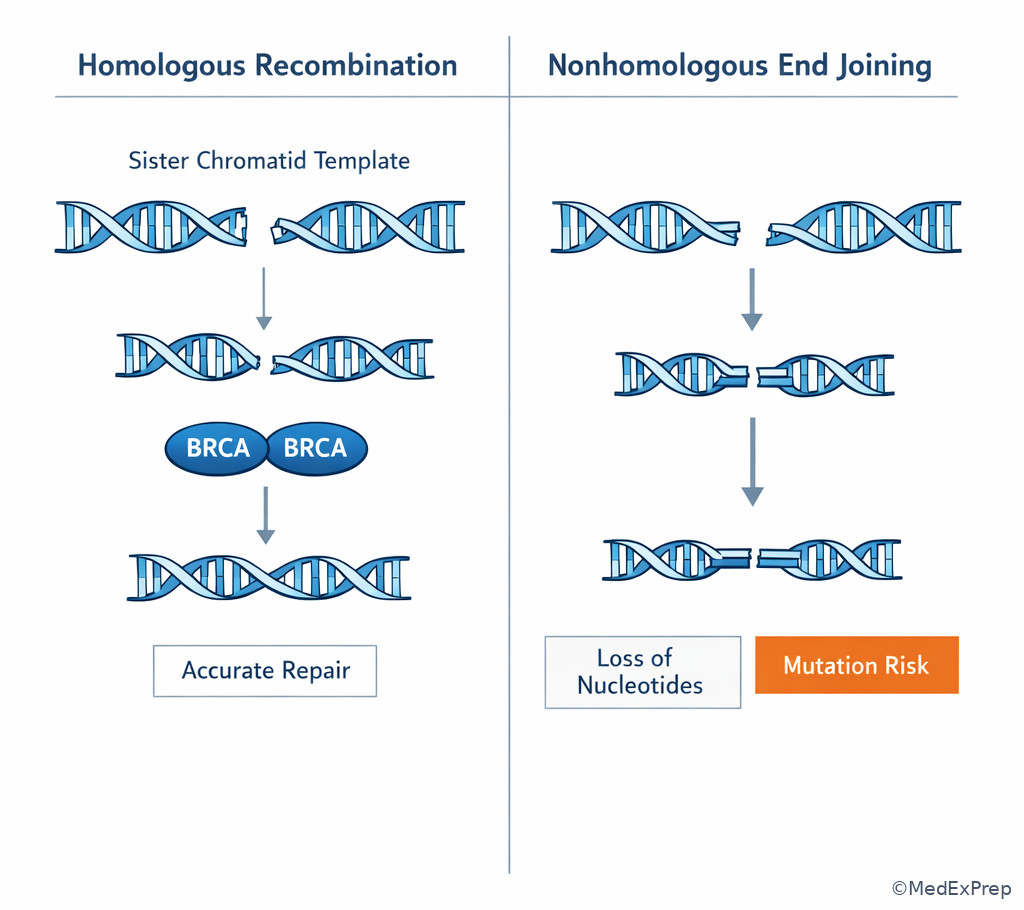
- Homologous recombination: High-fidelity repair using sister chromatid template
- Nonhomologous end joining: Double-strand break repair without template
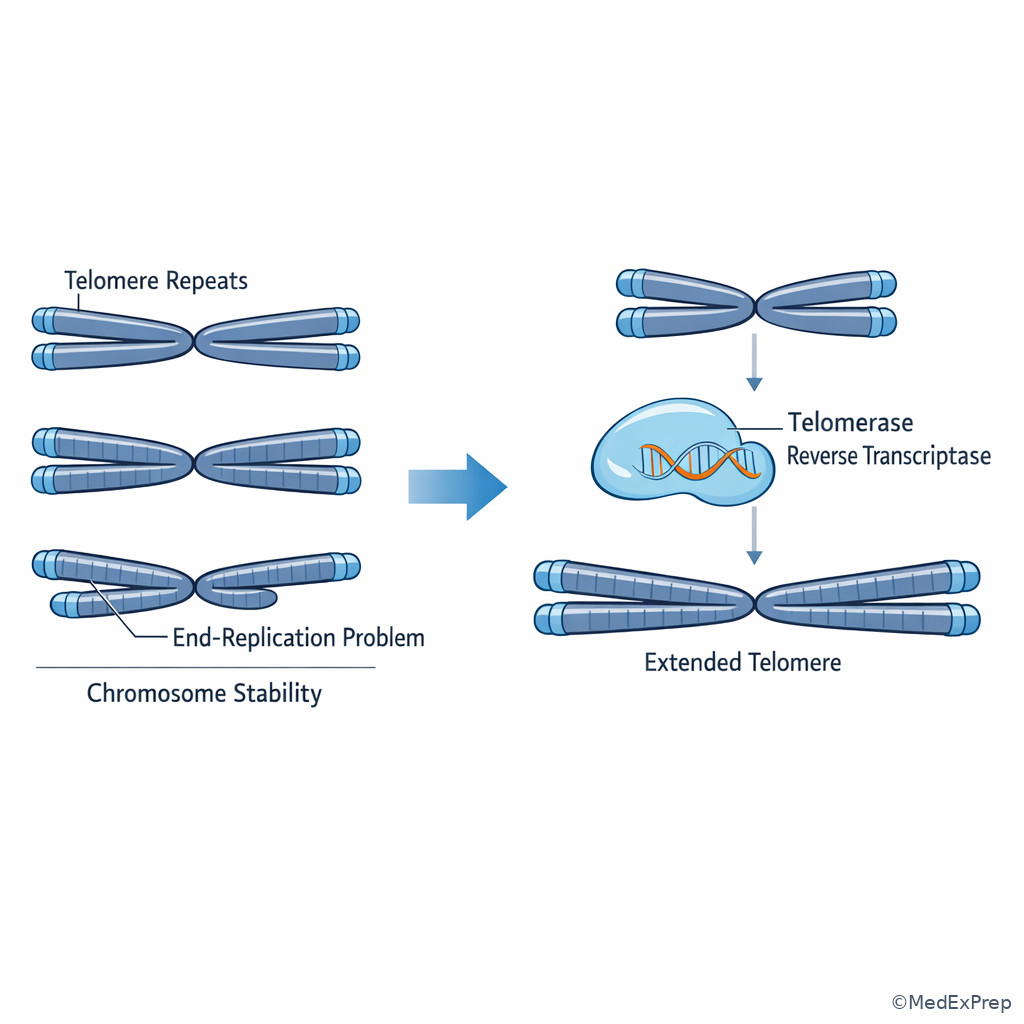
- Telomerase: Reverse transcriptase that extends chromosome ends
1.1.1.2 Must-know facts
- DNA polymerases synthesize only in the \(5' \to 3'\) direction.
- Reading of the template occurs in the \(3' \to 5'\) direction.
- Leading strand is continuous; lagging strand requires repeated priming.
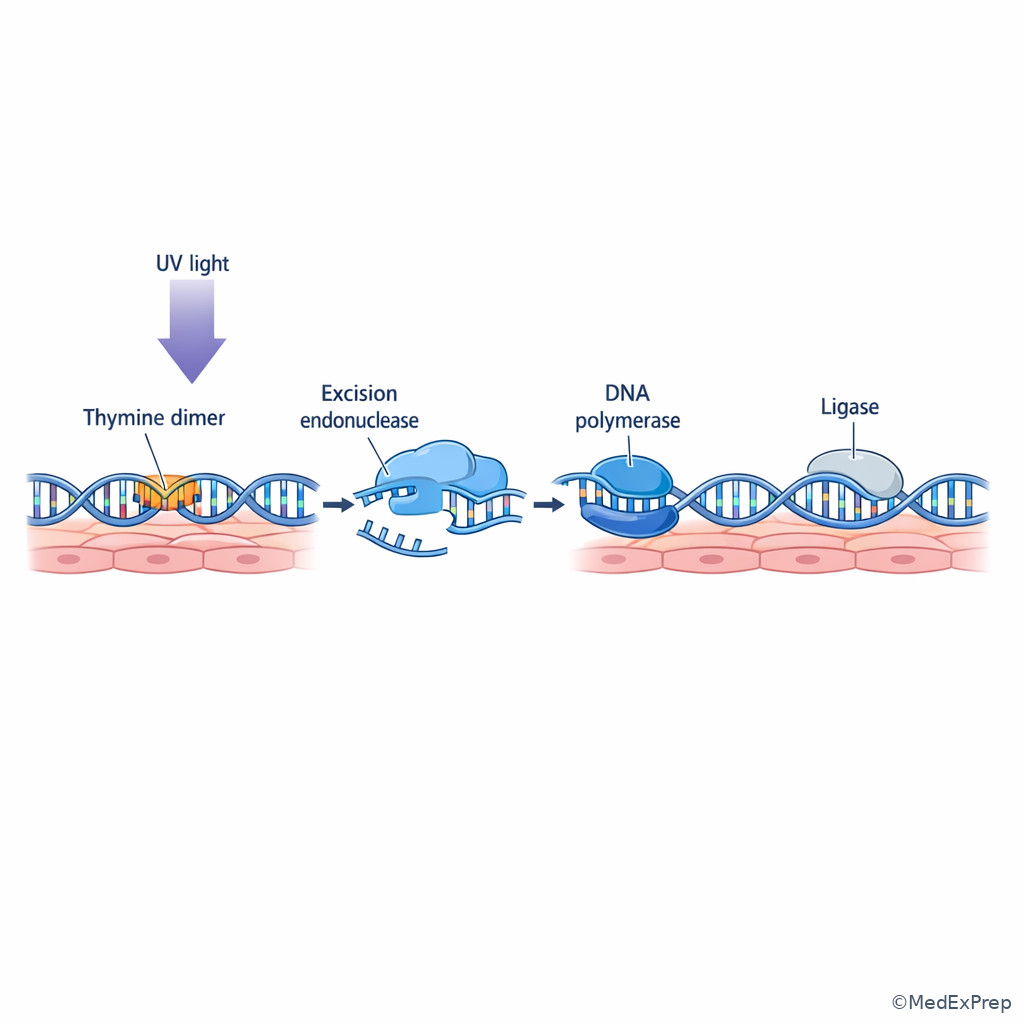
- UV light causes pyrimidine dimers, classically repaired by nucleotide excision repair.
- Depurination, deamination, and oxidation are usually handled by base excision repair.
- Lynch syndrome involves mismatch repair defects and microsatellite instability.
- Xeroderma pigmentosum involves nucleotide excision repair failure and marked UV sensitivity.
- Ataxia-telangiectasia involves defective double-strand break sensing/repair via ATM.
- BRCA1/BRCA2 are linked to homologous recombination repair of double-strand breaks.
- Telomere shortening limits replication in most somatic cells; telomerase is active in germ cells, stem cells, and many cancers.
Core content
Exam takeaway: Use fork direction, strand polarity, and primer dependence to separate helicase, primase, polymerase, ligase, and topoisomerase questions.
flowchart LR A[Origin opens] --> B[Helicase unwinds] B --> C[SSB stabilizes] C --> D[Primase lays RNA primer] D --> E[DNA polymerase extends 5 to 3] E --> F[Leading strand continuous] E --> G[Lagging strand Okazaki fragments] G --> H[Primer removal and replacement] H --> I[Ligase seals nick] B --> J[Topoisomerase relieves supercoils ahead of fork]
Exam takeaway: Match the type of lesion to the repair system rather than memorizing disease names alone.
flowchart TD
A[DNA damage found] --> B{What type?}
B -->|Replication mismatch| C[Mismatch repair]
B -->|Small base change or abasic site| D[Base excision repair]
B -->|Bulky adduct or pyrimidine dimer| E[Nucleotide excision repair]
B -->|Double-strand break| F{Template available?}
F -->|Yes sister chromatid| G[Homologous recombination]
F -->|No or rapid repair| H[Nonhomologous end joining] Exam takeaway: Visualize where each enzyme sits at the fork to avoid mixing up lagging-strand and proofreading questions.
Exam takeaway: Photosensitivity plus defective removal of bulky helix distortion should trigger nucleotide excision repair.
Exam takeaway: Homologous recombination uses a sister template for accuracy, whereas nonhomologous end joining trades speed for mutation risk.
Exam takeaway: Repeated replication shortens chromosome ends unless telomerase extends them, a key cancer and stem-cell concept.
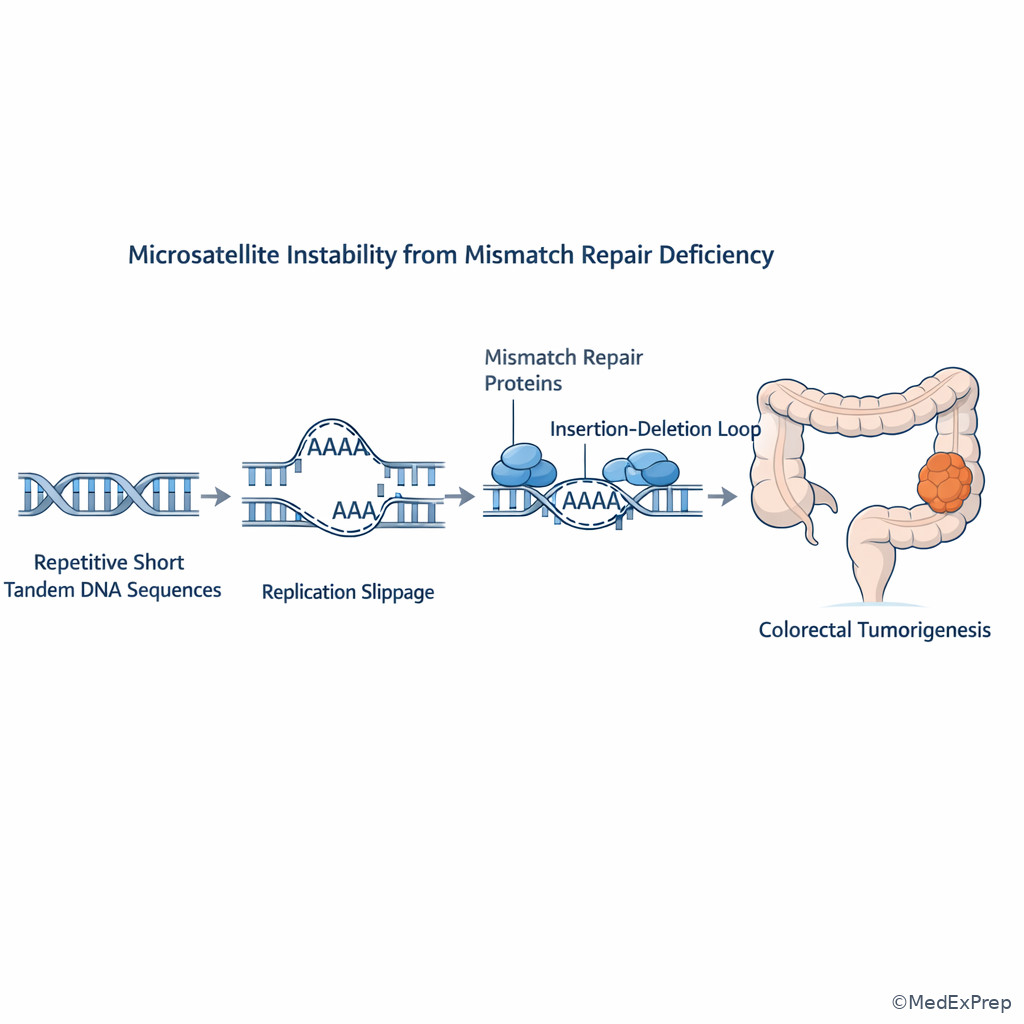
Exam takeaway: Repeated short DNA sequences become unstable when post-replication mismatch correction fails.
Exam takeaway: Small non-bulky base injury is removed one base at a time, not by wide excision of a DNA segment.
Exam takeaway: Defects in proofreading, mismatch repair, or double-strand break repair raise mutation burden through different mechanisms.
flowchart LR
A[Replication error] --> B{Corrected?}
B -->|Polymerase proofreading| C[Low mutation rate]
B -->|Missed then MMR works| C
B -->|MMR fails| D[Microsatellite instability]
A --> E[Double-strand break]
E --> F{Repair pathway}
F -->|Homologous recombination| G[Accurate repair]
F -->|Defective BRCA pathway| H[Chromosomal instability]
D --> I[Cancer predisposition]
H --> I Exam takeaway: UV, oxidation, deamination, and ionizing radiation create different lesions and therefore trigger different repair answers.
flowchart TD A[Damage source] --> B[UV light] A --> C[Reactive oxygen species] A --> D[Spontaneous deamination] A --> E[Ionizing radiation] B --> F[Pyrimidine dimers] C --> G[Oxidized bases] D --> H[Uracil in DNA] E --> I[Double-strand breaks] F --> J[Nucleotide excision repair] G --> K[Base excision repair] H --> K I --> L[Homologous recombination or NHEJ]
1.1.1.3 Foundational replication logic: what the fork is actually doing
A Explanation
Board questions here usually force a choice between enzymes that all sound “necessary for replication.” The decision conflict is simple: which exact physical problem at the fork must be solved? If the stem describes unwinding, choose helicase. If it describes torsional stress ahead of the fork, choose topoisomerase. If it describes starting synthesis, choose primase. If it describes adding DNA nucleotides, choose DNA polymerase. If it describes sealing fragments, choose ligase. See Fig 1 and Fig 3.
The single rule that resolves most items is polarity: DNA polymerases can only add nucleotides to a free 3' hydroxyl, so all new DNA is synthesized in the \(5' \to 3'\) direction. Because the two parental strands are antiparallel, the leading strand can be synthesized continuously toward the fork, whereas the lagging strand must be synthesized away from the fork in short Okazaki fragments. Each fragment needs a new RNA primer. Later, primer removal, gap filling, and ligation produce one intact daughter strand.
WHY THIS IS TESTED: students often memorize the enzyme list but cannot map an enzyme to the exact mechanical problem in the stem. COMLEX-style vignettes reward process reasoning, not list recall.
B Worked example
A 7-year-old boy is enrolled in a research protocol examining rapidly dividing bone marrow cells. He has asthma and eczema, and his mother mentions that he recently completed a course of amoxicillin for otitis media. Investigators add a toxin that prevents initiation of new nucleic acid strands but does not block elongation once synthesis has begun. Replication stalls most prominently on the lagging strand. What enzyme activity is most directly inhibited?
Reasoning chain: Identify task -> this is an enzyme-at-the-fork question. Extract key facts -> initiation blocked, elongation preserved, lagging strand especially affected. Apply rule -> lagging strand needs repeated starts, so repeated RNA primers are required. Eliminate distractors -> helicase would block unwinding of both strands; ligase failure would leave fragments unsealed after synthesis; topoisomerase failure causes torsional stress rather than failed initiation. Verify -> primase is the best answer because it creates the RNA primers needed to initiate each Okazaki fragment.
C Exam trap
D Checkpoint
Question: A toxin causes accumulation of positive supercoils ahead of a replication fork in human cells. Which enzyme is most directly impaired?
- Helicase — tempting because fork opening depends on it, but helicase separates strands at the fork rather than relieving torsional strain ahead of it.
- Topoisomerase — correct because this enzyme prevents overwinding created by strand separation during replication.
- Primase — tempting because replication requires repeated priming, but primer synthesis does not resolve supercoils.
- Ligase — tempting because it is essential for DNA continuity, but it seals backbone nicks after synthesis rather than reducing torsional stress.
1.1.1.4 Fidelity and proofreading: why replication is usually accurate
A Explanation
The next decision conflict is whether an error is corrected during synthesis or after synthesis. Proofreading is an intrinsic polymerase function that removes a mispaired nucleotide immediately using \(3' \to 5'\) exonuclease activity. Mismatch repair acts later, after replication, when the newly synthesized strand contains a base mismatch or small insertion-deletion loop missed by proofreading.
The single rule: if the vignette describes a polymerase detecting and removing the just-added wrong nucleotide at the growing 3' end, that is proofreading. If the stem describes post-replication correction, especially in repetitive DNA with microsatellite instability, that is mismatch repair. See Fig 2.
High-yield disease link: inherited mismatch repair defects cause Lynch syndrome, which is associated with colorectal and endometrial cancers and microsatellite instability. The trap is to confuse this with familial adenomatous polyposis, which involves APC mutation and numerous polyps rather than mismatch repair failure.
WHY THIS IS TESTED: board questions often pair “proofreading defect” and “mismatch repair defect” because both increase mutation burden, but they are not the same level of quality control.
B Worked example
A 42-year-old woman undergoes evaluation after her father had colon cancer at age 46 and her sister had endometrial cancer at age 39. She has type 2 diabetes and migraine headaches, and her laboratory panel incidentally shows mild isolated hypertriglyceridemia. Tumor testing demonstrates instability in repetitive short tandem DNA sequences. Which DNA repair process is most likely defective?
Reasoning chain: Identify task -> inherited cancer syndrome mechanism. Extract key facts -> early colon and endometrial cancers, microsatellite instability. Apply rule -> microsatellite instability points to mismatch repair failure. Eliminate distractors -> nucleotide excision repair gives UV sensitivity; base excision repair handles single abnormal bases; homologous recombination defects classically suggest BRCA-associated breast/ovarian syndromes. Verify -> mismatch repair is the single best answer.
C Exam trap
D Checkpoint
Question: A mutation abolishes the \(3' \to 5'\) exonuclease activity of a DNA polymerase but leaves nucleotide addition intact. Which function is lost?
- RNA primer synthesis — tempting because initiation is essential, but this is the role of primase, not polymerase proofreading.
- Proofreading of the newly added nucleotide — correct because \(3' \to 5'\) exonuclease activity removes misincorporated bases during synthesis.
- Removal of bulky thymine dimers — tempting because repair is involved, but bulky lesions are handled by nucleotide excision repair.
- Sealing of Okazaki fragment nicks — tempting because both involve lagging-strand completion, but ligase performs nick sealing.
- The most common repair pathway tested with microsatellite instability is mismatch repair.
- True or False: DNA polymerase synthesizes new DNA in the \(3' \to 5'\) direction. False.
- Name the 3 key lagging-strand features: repeated priming, Okazaki fragments, ligation.
1.1.1.5 Base excision versus nucleotide excision repair: choose the right lesion
A Explanation
This is one of the most testable comparisons in the lesson. The decision conflict is not “which repair pathway fixes DNA?” but rather which repair pathway fixes this exact kind of DNA damage? Small, non-bulky single-base lesions such as deamination, depurination, and oxidation are handled by base excision repair. Bulky helix-distorting lesions such as thymine dimers from UV light are handled by nucleotide excision repair. See Fig 2 and Fig 4.
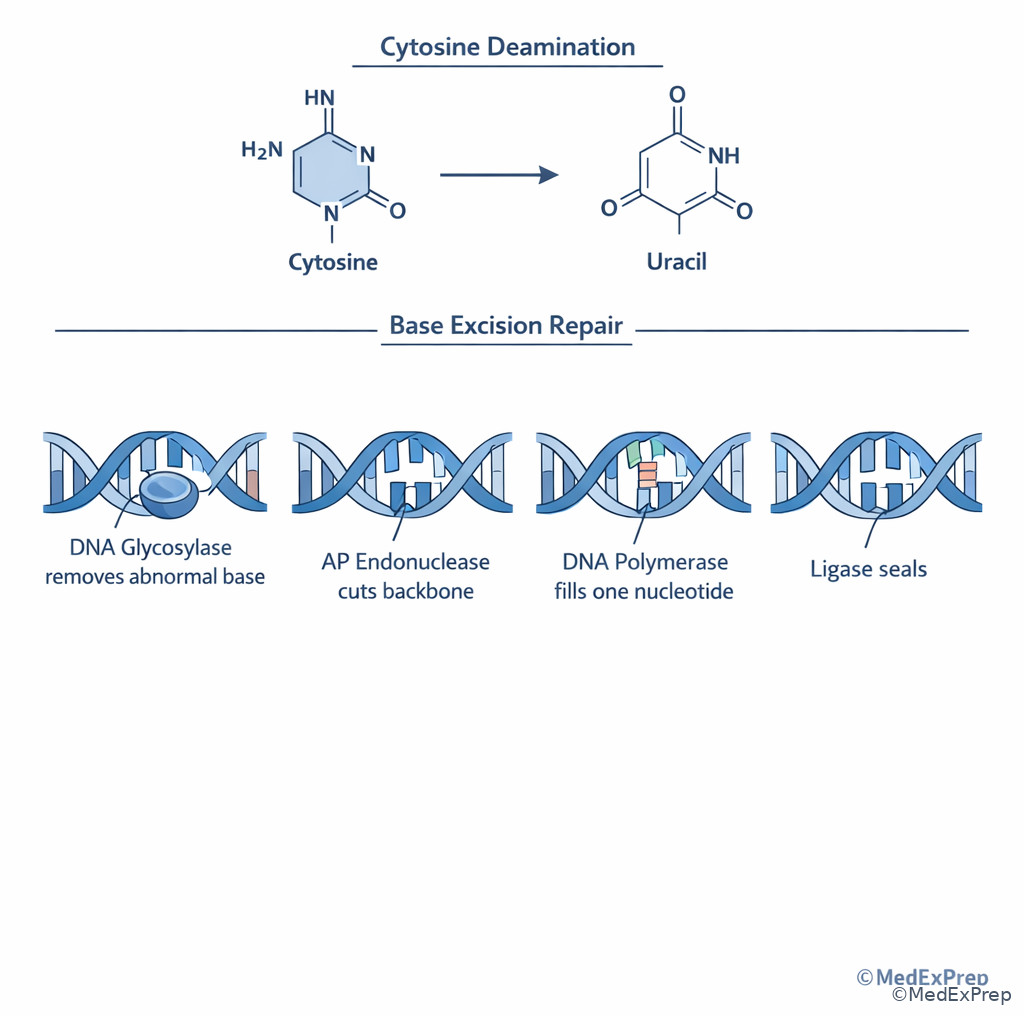
Base excision repair starts with a DNA glycosylase that removes the abnormal base, creating an abasic site. AP endonuclease cuts the backbone, then polymerase fills the missing nucleotide and ligase seals the nick. Nucleotide excision repair instead removes a short stretch of nucleotides surrounding the bulky lesion because the distortion affects more than a single base.
The single rule: small base change = base excision; bulky distortion = nucleotide excision. Xeroderma pigmentosum is the classic disease for defective nucleotide excision repair, producing extreme photosensitivity and skin cancer risk. A deaminated cytosine becoming uracil is the classic base excision setup.
WHY THIS IS TESTED: both pathways involve “cut, replace, seal,” so students confuse them unless they classify the lesion type first.
B Worked example
An 11-year-old boy is brought for evaluation of multiple freckle-like pigmented lesions on sun-exposed areas. He has a history of mild allergic rhinitis and repaired umbilical hernia, and his CBC shows a slightly low mean platelet volume that is not clinically relevant. His parents report severe sun sensitivity since early childhood. Biopsy of one lesion shows squamous cell carcinoma. Failure of which repair process best explains this presentation?
Reasoning chain: Identify task -> inherited DNA repair disorder. Extract key facts -> severe photosensitivity, sun-exposed lesions, early skin cancer. Apply rule -> UV causes pyrimidine dimers, repaired by nucleotide excision repair. Eliminate distractors -> base excision repair would fit deamination or oxidation; mismatch repair fits Lynch syndrome; homologous recombination fits BRCA-type double-strand break defects. Verify -> nucleotide excision repair failure is the answer, consistent with xeroderma pigmentosum.
C Exam trap
D Checkpoint
Question: Spontaneous deamination converts a cytosine to uracil within DNA. Which repair pathway most directly corrects this lesion?
- Mismatch repair — tempting because an incorrect base is present, but this pathway mainly corrects replication mismatches after DNA synthesis.
- Base excision repair — correct because a single abnormal non-bulky base is removed by glycosylase-mediated repair.
- Nucleotide excision repair — tempting because excision occurs here too, but this pathway is for bulky helix-distorting lesions such as thymine dimers.
- Nonhomologous end joining — tempting if any “DNA repair” option seems broad enough, but it is specific for double-strand breaks.
1.1.1.6 Double-strand break repair and cancer predisposition
A Explanation
Double-strand breaks create the highest-stakes DNA damage in this lesson. The decision conflict is whether the cell can perform accurate template-guided repair or must use a faster but error-prone method. If a sister chromatid is available, homologous recombination repairs the break with high fidelity. If no template is available or the cell uses rapid ligation, nonhomologous end joining directly rejoins broken ends and may lose nucleotides, increasing mutation risk. See Fig 5.
High-yield disease links: BRCA1 and BRCA2 participate in homologous recombination repair. When defective, cells accumulate chromosomal instability and cancer risk, especially breast and ovarian malignancies. ATM is a kinase involved in sensing double-strand DNA damage and coordinating repair; its deficiency causes ataxia-telangiectasia, classically associated with cerebellar ataxia, telangiectasias, sinopulmonary infections, and increased malignancy risk.
The single rule: if the clue is inherited breast/ovarian cancer susceptibility, think homologous recombination and BRCA. If the clue is radiation sensitivity with neurologic findings and telangiectasias, think ATM and defective double-strand break response. This is also where ionizing radiation belongs, because it classically causes double-strand breaks.
WHY THIS IS TESTED: several famous syndromes involve “DNA repair,” but the board expects you to separate UV lesions, mismatch errors, and double-strand breaks by the stem’s signature clue.
B Worked example
A 19-year-old man has progressive gait instability and recurrent sinus infections. Physical examination shows conjunctival telangiectasias. He also has seasonal allergies and recently sprained his ankle playing basketball. Lymphocyte culture demonstrates marked sensitivity to ionizing radiation with poor response to double-strand DNA injury. Defect in which protein best explains this finding?
Reasoning chain: Identify task -> syndrome linked to DNA repair. Extract key facts -> ataxia, telangiectasias, recurrent infections, radiation sensitivity. Apply rule -> classic ataxia-telangiectasia due to ATM defect, involving sensing/repair coordination for double-strand breaks. Eliminate distractors -> BRCA defects suggest hereditary breast/ovarian cancers; mismatch repair causes Lynch syndrome; nucleotide excision repair causes xeroderma pigmentosum. Verify -> ATM is correct.
C Exam trap
D Checkpoint
Question: A tumor cell line lacks BRCA1 function. Which DNA repair mechanism is most directly impaired?
- Homologous recombination repair of double-strand breaks — correct because BRCA proteins help accurate template-guided repair using a sister chromatid.
- Mismatch repair of replication errors — tempting because both defects increase cancer risk, but microsatellite instability rather than BRCA loss points to mismatch repair.
- Nucleotide excision repair of UV-induced dimers — tempting because cancer predisposition can result, but UV-sensitive disorders classically involve excision endonuclease pathways.
- Base excision repair of oxidized bases — tempting because single-base damage is common, but BRCA is not the key pathway for this lesion type.
- The most classic lesion from ionizing radiation is double-strand DNA break.
- True or False: Nonhomologous end joining is more accurate than homologous recombination. False.
- Name the 3 classic clues for ataxia-telangiectasia: ataxia, telangiectasias, radiation sensitivity.
1.1.1.7 Telomeres, end-replication problem, and cellular immortality
A Explanation
The final foundational concept asks what happens at chromosome ends. The decision conflict is whether a cell can fully replicate linear DNA termini indefinitely. Because DNA polymerase requires a primer and can only extend from an existing 3' OH, removal of the terminal RNA primer on the lagging strand leaves a gap that cannot be filled conventionally. This is the end-replication problem. Repeated cycles therefore shorten telomeres in most somatic cells. See Fig 6.
Telomerase is a reverse transcriptase that carries its own RNA template and extends telomeric repeats, preserving chromosome ends. It is active in germ cells, stem cells, and many cancers. The single rule: if the question asks what allows malignant cells to bypass replicative senescence, think telomerase reactivation. If it asks why normal somatic cells have finite replicative capacity, think progressive telomere shortening.
This topic also connects to aging, oncogenesis, and hematopoietic stem cell biology. COMLEX may frame it as a mechanism question rather than as pure cell biology. The answer is usually not “DNA polymerase becomes stronger”; it is specific reactivation of telomerase or altered checkpoint control allowing immortalization.
WHY THIS IS TESTED: telomerase is an easy concept to memorize superficially, but exam stems often hide it inside a cancer or stem-cell scenario.
B Worked example
A 63-year-old woman with obesity and hypertension has a resected colon adenocarcinoma used to create a laboratory cell line. She previously had cholecystectomy and has a mildly elevated alkaline phosphatase from osteoarthritis-related bone turnover. The cell line continues dividing far beyond the usual number of passages seen in normal fibroblasts. Which molecular change best explains this persistent proliferative capacity?
Reasoning chain: Identify task -> mechanism of unlimited cell division. Extract key facts -> cancer cell line, repeated passage, continued proliferation. Apply rule -> immortalization is classically linked to telomerase activation. Eliminate distractors -> increased ligase or primase would not solve chromosome-end shortening; mismatch repair loss increases mutation rate but does not itself directly create indefinite replicative capacity. Verify -> telomerase reactivation is the answer.
C Exam trap
D Checkpoint
Question: Which cell type normally has high telomerase activity?
- Mature erythrocyte — tempting because blood cells are renewed, but mature erythrocytes lack nuclei and do not replicate DNA.
- Skeletal muscle fiber — tempting because it is long-lived, but it is not a classic highly proliferative telomerase-rich cell type.
- Germ cell — correct because germ cells must preserve chromosome ends across repeated divisions.
- Most differentiated somatic epithelial cells — tempting because they divide more than neurons, but telomerase is still limited in most normal somatic cells.
Exam Traps & Differentiators
Most common wrong answer and why: choosing the broad category “DNA repair” without classifying lesion type first. The stem almost always gives one discriminating clue: UV exposure, microsatellite instability, ionizing radiation, or immediate proofreading at the fork.
| Looks similar | But isn’t | Single clue that separates them | Think |
|---|---|---|---|
| Proofreading defect | Mismatch repair defect | Error removed during synthesis vs post-replication microsatellite instability | Polymerase exonuclease vs Lynch syndrome |
| Base excision repair | Nucleotide excision repair | Small single-base lesion vs bulky UV-induced helix distortion | Deamination/oxidation vs thymine dimers |
| BRCA-related repair defect | ATM-related repair defect | Breast/ovarian cancer predisposition vs ataxia-telangiectasia with radiation sensitivity | Homologous recombination vs double-strand break sensing |
| Lagging-strand primase problem | Ligase problem | Failure to start fragments vs failure to seal completed fragments | Initiation vs completion |
| Telomerase activation | Increased polymerase speed | Unlimited cell division due to chromosome-end maintenance | Immortalization mechanism |
- If the stem says “severe photosensitivity with early skin cancers,” think nucleotide excision repair failure, not base excision repair.
- If the stem says “microsatellite instability” or “colon plus endometrial cancers,” think mismatch repair.
- If the stem says “ataxia, telangiectasias, recurrent infections, radiation sensitivity,” think ATM and double-strand break response.
- If the stem says “error removed from the newly added 3' end,” think polymerase proofreading.
- If the stem says “repeated need to start lagging-strand synthesis,” think primase.
- If the stem says “cell line divides indefinitely,” think telomerase reactivation.
Tables
| Enzyme / pathway | Main job | Classic board clue | Best discriminator |
|---|---|---|---|
| Helicase | Separates parental DNA strands | Fork unwinding blocked | Problem occurs at strand separation itself |
| Topoisomerase | Relieves supercoiling ahead of fork | Torsional strain or positive supercoils | Acts ahead of fork, not at primer or nick |
| Primase | Makes RNA primer | Initiation defect, especially lagging strand | Needed before polymerase can extend |
| DNA polymerase | Extends DNA \(5' \to 3'\) | Elongation defect | Adds to free 3' OH only |
| Proofreading | Removes just-added wrong nucleotide | \(3' \to 5'\) exonuclease loss | During synthesis, not after replication |
| Ligase | Seals phosphodiester nicks | Unjoined Okazaki fragments | Completion step after synthesis |
| Telomerase | Extends telomeres | Immortalized cells | Solves end-replication problem |
| Comparison | Base excision repair | Nucleotide excision repair | Mismatch repair |
|---|---|---|---|
| When used | Small non-bulky base injury | Bulky helix-distorting lesion | Post-replication base mismatch or insertion-deletion loop |
| Typical lesions | Deamination, depurination, oxidation | Pyrimidine dimers, bulky adducts | Mispaired bases, microsatellite slippage |
| How much DNA removed | Usually one abnormal base | Short oligonucleotide segment | Patch around mismatch on daughter strand |
| Classic disease link | Less often tested as a named syndrome | Xeroderma pigmentosum | Lynch syndrome |
| Signature clue | Uracil in DNA | Photosensitivity | Microsatellite instability |
| Differential / classification | Defect | Clinical pattern | High-yield clue |
|---|---|---|---|
| Xeroderma pigmentosum | Nucleotide excision repair | Extreme UV sensitivity, early skin cancers | Pyrimidine dimer removal failure |
| Lynch syndrome | Mismatch repair | Colon and endometrial cancer predisposition | Microsatellite instability |
| BRCA-associated hereditary cancer | Homologous recombination | Breast/ovarian cancer susceptibility | Double-strand break repair defect |
| Ataxia-telangiectasia | ATM-mediated double-strand break response | Ataxia, telangiectasias, infections | Radiation sensitivity |
| Familial adenomatous polyposis | APC tumor suppressor mutation | Numerous colonic polyps | Not a mismatch repair syndrome |
| Damage source | Main lesion | Most likely repair answer | Common wrong answer |
|---|---|---|---|
| UV light | Pyrimidine dimers | Nucleotide excision repair | Base excision repair |
| Spontaneous deamination | Uracil in DNA | Base excision repair | Nucleotide excision repair |
| Reactive oxygen species | Oxidized base | Base excision repair | Mismatch repair |
| Replication slippage | Insertion-deletion loop | Mismatch repair | Proofreading |
| Ionizing radiation | Double-strand break | Homologous recombination or NHEJ | Nucleotide excision repair |
Algorithm / Approach
Use this mini-algorithm whenever the answer choices are all DNA enzymes or all repair pathways. First decide whether the issue is fork mechanics or damage response. Then classify the exact defect. That two-step approach prevents most near-miss errors.
Rapid Review
- Helicase → separates strands at the fork, not supercoils ahead of it
- Topoisomerase → relieves torsional stress ahead of fork, classic chemotherapy target
- Primase → required repeatedly on lagging strand because each Okazaki fragment needs a new start
- DNA polymerase → extends only from a free 3' OH in the \(5' \to 3'\) direction
- Proofreading → immediate correction of just-added nucleotide via \(3' \to 5'\) exonuclease
- Mismatch repair → post-replication error correction, classic microsatellite instability clue
- Base excision repair → single abnormal base removed, think deamination or oxidation
- Nucleotide excision repair → bulky helix distortion removed, think UV pyrimidine dimers
- Xeroderma pigmentosum → photosensitivity plus skin cancers from NER failure
- Lynch syndrome → colon and endometrial cancer pattern from mismatch repair loss
- Homologous recombination → accurate double-strand break repair using sister chromatid
- Nonhomologous end joining → quick but error-prone break repair without template
- BRCA1/2 → homologous recombination, not mismatch repair
- ATM → radiation sensitivity with ataxia-telangiectasia phenotype
- Telomerase → reverse transcriptase that supports cellular immortality in many cancers
Self-check quiz
1.1.1.8 Question 1
A 22-year-old woman in a molecular biology lab is asked which enzyme relieves positive supercoils generated ahead of a moving replication fork. She has mild iron deficiency and seasonal allergies. Which is the most appropriate answer?
- Helicase
- Topoisomerase
- Primase
- Ligase
1.1.1.9 Question 2
A 9-year-old boy with severe sun sensitivity develops multiple pigmented lesions on sun-exposed skin. He also has mild eczema and a recent history of streptococcal pharyngitis. Defect in which pathway most likely explains this presentation?
- Base excision repair
- Mismatch repair
- Nucleotide excision repair
- Nonhomologous end joining
1.1.1.10 Question 3
A 45-year-old woman undergoes evaluation because her mother had colon cancer at 44 and her aunt had endometrial cancer at 48. She has obesity and hypertension, and her lipid panel shows mild hypertriglyceridemia. Tumor testing reveals microsatellite instability. Which mechanism is most likely defective?
- Repair of bulky thymine dimers after UV exposure
- Post-replication correction of base mismatches and insertion-deletion loops
- Direct ligation of broken DNA ends without a template
- Extension of chromosome ends by reverse transcriptase
1.1.1.11 Question 4
A 6-year-old child is exposed in a laboratory accident to an agent that prevents synthesis of short RNA segments needed to start DNA replication. He has asthma and repaired ventricular septal defect, and a chest radiograph obtained last month for cough was normal. Which process will be most immediately impaired?
- Relief of torsional strain ahead of the fork
- Initiation of Okazaki fragment synthesis on the lagging strand
- Sealing of phosphodiester nicks after elongation
- Excision of UV-induced pyrimidine dimers
1.1.1.12 Question 5
A 28-year-old man with no major medical history participates in a radiation biology study. After ionizing radiation exposure, his cultured cells show a selective inability to perform high-fidelity repair using a sister chromatid template. Which pathway is most directly impaired?
- Base excision repair
- Homologous recombination
- Mismatch repair
- Nucleotide excision repair
1.1.1.13 Question 6
A 17-year-old boy has progressive gait instability, recurrent sinopulmonary infections, and conjunctival telangiectasias. He also has allergic rhinitis and an elevated LDL cholesterol level found on screening. Cells obtained from the patient are unusually sensitive to ionizing radiation. Which defect best explains his condition?
- ATM-mediated response to double-strand DNA breaks
- BRCA1-mediated homologous recombination in breast epithelium
- Mismatch repair of repetitive DNA sequences
- Nucleotide excision repair of UV-induced dimers
1.1.1.14 Question 7
A 64-year-old woman with a resected adenocarcinoma has a tumor cell line established for research. She also has type 2 diabetes and osteoarthritis, and her alkaline phosphatase is mildly elevated. The cell line continues dividing after many passages, unlike matched normal fibroblasts. Which change most likely permits this persistent proliferation?
- Loss of nucleotide excision repair
- Increased telomerase activity
- Loss of RNA primer removal
- Increased DNA ligase activity
1.1.1.15 Question 8
A researcher introduces a mutation that abolishes exonuclease activity responsible for removing incorrectly paired nucleotides immediately after they are added to the growing DNA strand. The cells have normal mismatch repair proteins, normal ligase, and normal UV-damage repair. Which defect is most directly present?
- Failure of proofreading during DNA synthesis
- Failure of post-replication mismatch repair
- Failure of nucleotide excision repair
- Failure of nonhomologous end joining
Answer key
1.1.1.16 Question 1: B. Topoisomerase
Correct-answer mechanism: Unwinding of DNA by helicase creates torsional stress ahead of the replication fork. Topoisomerase transiently cuts and rejoins DNA to relieve positive supercoiling, allowing fork progression. This is the classic “supercoils ahead of the fork” clue. Sources: Alberts, Molecular Biology of the Cell; First Aid for the USMLE Step 1.
- A. Helicase — tempting because it is essential for strand separation, but the single discriminating clue is ahead of the fork; helicase acts at the fork itself, not on supercoils ahead of it.
- B. Topoisomerase — tempting and correct because it directly resolves overwinding generated during replication; the key clue is positive supercoils.
- C. Primase — tempting because all replication requires it, but primase lays RNA primers and does not alter DNA topology.
- D. Ligase — tempting because it is central to lagging-strand processing, but it seals nicks after synthesis rather than relieving torsional strain.
1.1.1.17 Question 2: C. Nucleotide excision repair
Correct-answer mechanism: Severe UV sensitivity with early skin lesions strongly suggests xeroderma pigmentosum, caused by defective nucleotide excision repair. UV light creates pyrimidine dimers, which are bulky helix-distorting lesions removed by excision of a short DNA segment around the lesion. Sources: Robbins & Cotran; First Aid for the USMLE Step 1.
- A. Base excision repair — tempting because it also removes damaged DNA, but the discriminating clue is sun sensitivity; BER handles small single-base lesions, not bulky UV dimers.
- B. Mismatch repair — tempting because repair defects predispose to cancer, but the clue for MMR would be microsatellite instability and Lynch-type tumors, not photosensitivity.
- C. Nucleotide excision repair — correct because UV-induced pyrimidine dimers distort the helix and require removal of an oligonucleotide patch.
- D. Nonhomologous end joining — tempting if any DNA-repair label seems broad enough, but NHEJ is for double-strand breaks, especially when no homologous template is used.
1.1.1.18 Question 3: B. Post-replication correction of base mismatches and insertion-deletion loops
Correct-answer mechanism: Microsatellite instability in a family with early colon and endometrial cancers points to Lynch syndrome due to mismatch repair deficiency. Mismatch repair corrects errors missed by proofreading after DNA replication is completed, especially insertion-deletion loops in repetitive sequences. Sources: Harrison’s; Robbins & Cotran.
- A. Repair of bulky thymine dimers after UV exposure — tempting because it is another classic repair pathway, but the discriminating clue is microsatellite instability, not photosensitivity.
- B. Post-replication correction of base mismatches and insertion-deletion loops — correct because this is the core function of mismatch repair, the pathway lost in Lynch syndrome.
- C. Direct ligation of broken DNA ends without a template — tempting because cancer can result from genomic instability, but NHEJ defects are not the classic cause of microsatellite instability.
- D. Extension of chromosome ends by reverse transcriptase — tempting because cancer cells often use telomerase, but the family-history clue points specifically to hereditary mismatch repair deficiency.
1.1.1.19 Question 4: B. Initiation of Okazaki fragment synthesis on the lagging strand
Correct-answer mechanism: Short RNA segments needed to start replication are RNA primers made by primase. Lagging-strand synthesis is most immediately affected because each Okazaki fragment requires a new primer, whereas the leading strand needs only one initial primer. Sources: Alberts; First Aid for the USMLE Step 1.
- A. Relief of torsional strain ahead of the fork — tempting because fork progression depends on it, but that is topoisomerase function, not primer synthesis.
- B. Initiation of Okazaki fragment synthesis on the lagging strand — correct because loss of RNA primer synthesis prevents repeated starts on the lagging strand.
- C. Sealing of phosphodiester nicks after elongation — tempting because lagging-strand maturation also needs ligase, but the clue says start DNA replication, not finish it.
- D. Excision of UV-induced pyrimidine dimers — tempting as another nucleic-acid pathway, but this is a DNA repair function unrelated to primer synthesis.
1.1.1.20 Question 5: B. Homologous recombination
Correct-answer mechanism: High-fidelity repair using a sister chromatid template is homologous recombination. This pathway is critical for accurate repair of double-strand breaks and is associated with BRCA proteins. Sources: Robbins & Cotran; Harrison’s.
- A. Base excision repair — tempting because radiation can generate reactive species, but the discriminating clue is sister chromatid template, which points to double-strand break repair via homologous recombination.
- B. Homologous recombination — correct because it uniquely uses homologous DNA as a template for accurate repair.
- C. Mismatch repair — tempting because replication-associated errors also need high fidelity, but MMR corrects mismatched bases, not double-strand breaks using a sister chromatid.
- D. Nucleotide excision repair — tempting because radiation and mutagens may broadly damage DNA, but NER addresses bulky lesions rather than template-guided double-strand break repair.
1.1.1.21 Question 6: A. ATM-mediated response to double-strand DNA breaks
Correct-answer mechanism: Ataxia, telangiectasias, recurrent sinopulmonary infections, and ionizing radiation sensitivity identify ataxia-telangiectasia due to ATM mutation. ATM is a kinase involved in sensing DNA double-strand breaks and coordinating checkpoint activation and repair. Sources: Robbins & Cotran; First Aid for the USMLE Step 1.
- A. ATM-mediated response to double-strand DNA breaks — correct because the combination of cerebellar findings, telangiectasias, immunodeficiency, and radiation sensitivity is classic for ATM deficiency.
- B. BRCA1-mediated homologous recombination in breast epithelium — tempting because BRCA also relates to double-strand break repair, but the discriminating clue is the syndrome phenotype, not hereditary breast/ovarian cancer.
- C. Mismatch repair of repetitive DNA sequences — tempting because inherited DNA repair defects cause cancer risk, but the key clue of microsatellite instability is absent.
- D. Nucleotide excision repair of UV-induced dimers — tempting because repair defects can cause cancer, but the discriminating clue would be photosensitivity and skin cancers, not neurologic and immune findings.
1.1.1.22 Question 7: B. Increased telomerase activity
Correct-answer mechanism: Persistent proliferation after many passages indicates bypass of replicative senescence. Cancer cells commonly reactivate telomerase, a reverse transcriptase that extends telomeric repeats and counteracts chromosome-end shortening. Sources: Harrison’s; First Aid for the USMLE Step 1.
- A. Loss of nucleotide excision repair — tempting because defective repair promotes carcinogenesis, but the discriminating clue is continued division over many passages, which specifically points to telomere maintenance.
- B. Increased telomerase activity — correct because it permits maintenance of chromosome ends and supports cellular immortality.
- C. Loss of RNA primer removal — tempting because it affects replication processing, but such a defect would impair DNA synthesis rather than enhance long-term proliferative capacity.
- D. Increased DNA ligase activity — tempting because ligase is needed to finish replication, but better nick sealing does not solve the end-replication problem.
1.1.1.23 Question 8: A. Failure of proofreading during DNA synthesis
Correct-answer mechanism: Immediate removal of an incorrectly paired nucleotide right after addition to the growing strand is proofreading, mediated by the polymerase’s \(3' \to 5'\) exonuclease activity. The stem explicitly preserves mismatch repair, so this is not a post-replication correction defect. Sources: Alberts; First Aid for the USMLE Step 1.
- A. Failure of proofreading during DNA synthesis — correct because the discriminating clue is immediately after they are added, which defines proofreading.
- B. Failure of post-replication mismatch repair — tempting because mismatch repair also handles incorrect base pairing, but the stem specifically says mismatch repair proteins are normal and describes an immediate correction step.
- C. Failure of nucleotide excision repair — tempting if “exonuclease” suggests excision generally, but NER addresses bulky lesions rather than removal of the just-added incorrect base.
- D. Failure of nonhomologous end joining — tempting only as another repair pathway, but the discriminating clue is not a double-strand break; it is a misinserted nucleotide at the replication fork.
